New research could help design nanocontainers for drug delivery
Posted: 11 March 2022 | Ria Kakkad (Drug Target Review) | No comments yet
New research by UC Riverside may be a crucial step in the design of engineered nano-shells that could serve as vehicles for delivering drugs to specific targets in the body.

An RNA virus’s genome dictates how the virus replicates in cells to eventually cause disease. The genome also has the code for making a capsid, the protein shell of a virus that encapsulates the genome and protects it like a nanocontainer. The University of California, Riverside, US has published a study in ACS Nano, which may help to explain how a virus finds its native genome and how capsids form around it and not around other RNAs in the cell.
“A better understanding of how capsids form is of vital importance to material scientists and a crucial step in the design of engineered nano-shells that could serve as vehicles for delivering drugs to specific targets in the body,” said Professor Roya Zandi, lead researcher.
The research showed that the interaction of the mechanical properties of proteins, the size of the genome and the strength of the interaction between the genome and capsid proteins can significantly modify the symmetry, structure and stability of the capsid. When a virus enters a cell, the capsid breaks open to release the genome, which then uses the cell’s reproductive machinery to replicate. The newly formed genomes begin to acquire their capsids, a process mainly driven by the attractive electrostatic interaction between the positive charges on capsid proteins and the negative charges on the genomes. However, how the virus selects and packages its native RNA inside the crowded environment of a host cell cytoplasm in the presence of many non-viral RNA and other polymers has remained a mystery.
The simulations run by the researchers highlighted that theoretically, capsid proteins could pick any non-viral genome to encapsulate. However the viral genome, Zandi highlighted, is best suited for capsid proteins to form a shell around due to an interplay of energies at the molecular level.
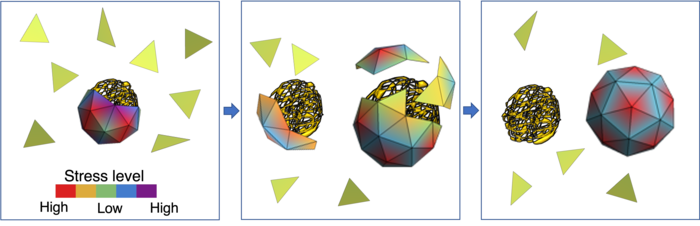

From left to right: The green triangles mimic capsid proteins. In the absence of a viral genome, they assemble around smaller segments of RNA. However, the shell is stressed and can be easily broken into pieces. The color bar shows the different levels of stress. Once the viral RNA is available, the capsid proteins start to leave non-viral RNAs and assemble around the viral genome. This can be done relatively easily because of the high stress level of capsid proteins in the smaller shell. The right figure shows capsid proteins assembled around the native RNA, forming a stable icosahedral shell.
[Credit: Zandi lab, UC Riverside.]
“The stress distribution of the capsid proteins is lower when the capsids encapsulate their own genome, the one for which they were coded,” Zandi said. “The energy of the whole system is lower. While smaller non-viral RNAs are available in the cell in plenty, the capsid proteins are inclined towards forming a shell around a viral RNA because the resulting soccer ball-like shell has a lower stress distribution.”
The researchers hope their study will allow for a better understanding of the role of RNA in the capsid assembly pathway, stability and structure.
Related topics
Drug Delivery, Drug Targets, Genome Editing, RNAs
Related organisations
University of California Riverside
Related people
Professor Roya Zandi