New control system created for synthetic genes
Posted: 3 November 2022 | Izzy Wood (Drug Target Review) | No comments yet
MIT researchers have developed a technique that could help the production of monoclonal antibodies and other useful proteins.

MIT researchers, using an approach based on CRISPR proteins, have developed a new way to precisely control the amount of a particular protein that is produced in mammalian cells.
This technique could be used to finely tune the production of useful proteins, such as the monoclonal antibodies used to treat cancer and other diseases. In their new study, which appeared in Nature Communications, the researchers showed that this system can work in a variety of mammalian cells, with very consistent results.
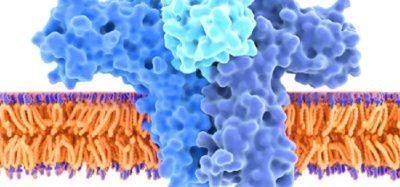
Many therapeutic proteins are produced in large bioreactors containing mammalian cells that are engineered to generate the desired protein. To do so, the researchers targeted the promoters of the genes they wanted to upregulate. In all mammalian cells, genes have a promoter region that binds to transcription factors.
In 2013, researchers in Lu’s lab developed a CRISPR-based transcription factor that allowed them to more easily control transcription of naturally occurring genes in mammalian and yeast cells.
In the new study, the researchers set out to build on that work, creating a library of synthetic biological parts that would allow them to deliver a transgene and precisely control its expression.
William Chen, assistant professor of biomedical sciences at the University of South Dakota, explained: “The idea is to have a full-spectrum synthetic promoter system that can go from very low to very high, to accommodate different cellular applications.”
The system designed includes several components:
- A gene to be transcribed, along with an “operator” sequence, which consists of a series of artificial transcription factor binding sites
- A guide RNA that binds to those operator sequences
- A transcription activation domain attached to a deactivated Cas9 protein. When this deactivated Cas9 protein binds to the guide RNA at the synthetic promoter site, the CRISPR-based transcription factor can turn on gene expression.
The promoter sites used for this synthetic system were designed to be distinct from naturally occurring promoter sites, so that the system will not affect genes in the cells’ own genomes.
Each operator includes between two and 16 copies of the guide RNA binding site and the researchers found that their system could initiate gene transcription at rates that linearly correspond to the number of binding sites, allowing them to precisely control the amount of the protein produced.
The system was tested in several types of mammalian cells, including Chinese hamster ovary (CHO) cells, which are commonly used to produce therapeutic proteins in industrial bioreactors. They found very similar results in CHO cells and the other cells they tested.
“The system has very high consistency over different cell types and different target genes,” Chen said. “This is a good starting point for thinking about regulating gene expression and cell behaviour with a highly tuneable, predictable artificial system.”
The researchers also programmed CHO cells to produce different quantities of a human antibody called anti-PD1. When human T cells were exposed to these cells, they became more potent tumour cell killers if there was a larger amount of the antibody produced.
Although the researchers were able to obtain a high yield of the desired antibodies, further work would be needed to incorporate this system into industrial processes, they say. Unlike the cells used in industrial bioreactors, the cells used in this study were grown on a flat surface, rather than in a liquid suspension.
“It is a highly predictable system that we can design up front and then get the expected outcome,” said William Chen, a former MIT research scientist. “It is a very tuneable system and suitable for many different biomedical applications in different cell types.”
Related topics
Drug Targets, Gene Testing, Gene Therapy, Genetic Analysis, Synthetic Biology, Targets, Therapeutics
Related organisations
MIT
Related people
William Chen