Creating the ‘google earth’ of tumours
Posted: 20 April 2017 | Josephine Bunch (Lead Scientist: NPL) | No comments yet
We caught up with Dr Josephine Bunch to find out how NPL are using the Cancer Research Grand Challenge funding & novel imaging techniques to map cancer tumours…

Scientists at NPL believe that ‘to improve chances of beating cancer, we need the ability to map and measure from the whole tumour scale, through tissue and cellular levels, all the way down to the molecular scale’. Now, thanks to the Cancer Research grand challenge, they have the funding to create this comprehensive map of tumours.
Back in February 2017, Cancer Research UK announced that four international teams are the first recipients of its global £100m Grand Challenge competition, which aims to overcome the biggest challenges facing cancer researchers in a global effort to beat cancer sooner. The winning projects are set to revolutionise our understanding of cancer, and enable us to better prevent, diagnose and treat the disease in the future. The international, multidisciplinary teams will be using unparalleled approaches to help develop pioneering solutions to some of cancer’s major challenges.
The Cancer Research Grand Challenge – how did you get involved? How will the money be spent and divided across your research project?
I first heard about the Grand Challenge on BBC Radio 4 in 2015 and immediately knew that the imaging techniques we were developing could provide a solution to the problem described by Cancer Research UK. The scale of the Grand Challenge funding allowed us to assemble an extremely ambitious programme of research that combines expertise in both the physical and life sciences in a way that’s never been done before. The award will be used to support new people, as well as to buy new instruments and equipment. We are appointing chemists, biologists, clinical fellows, physicists and computer scientists across the consortium and buying several new mass spectrometers. Our award will also support pre-clinical studies, analysis of clinical samples, and many other laboratory costs associated with the five-year programme.
Are you collaborating with any other institutions or companies and what will their input be?
Our multi-disciplinary team includes co-investigators from industry as well as research institutes:
- Prof Owen J Sansom, Cancer Research UK Beatson Institute
- Prof Zoltan Takats, Imperial College London
- Dr Richard Goodwin, AstraZeneca
- Dr George Poulogiannis, The Institute of Cancer Research
- Dr Simon Barry, AstraZeneca
- Dr Mariia Yuneva, The Francis Crick Institute
- Prof John Marshall, Barts Cancer Institute
- Prof Kevin Brindle, Cancer Research UK Cambridge Institute
- Prof Ian Gilmore, NPL
We also have two patient advocates in our consortium – Kelly Gleason and Harry C Hall.
What is meant by the term ‘Google Earth of tumours’?
The aim is to create a reproducible, standardised way to fully understand different tumours in unprecedented detail. To achieve this we’ll map every minutiae of several types of cancer; recording every aspect from intact tumour tissues to singles cells and the molecules held within them. From whole tumours right down to the individual fats and proteins in cells, we will map and visualise every element of these tumours to create faithful 3D representations for the first time. It’s the scale of imaging that makes us compare it to that of Google Earth – from providing an overall view of the whole tumour, right the way down to its molecular level. It can be compared to not only identifying an individual ‘house’ and where it is in a country, but also who’s inside the house, what they’re eating and watching on TV.
Which types of cancer will you be analysing and why?
The study will encompass individual breast, bowel and pancreatic tumours. We are working with world experts in cancer metabolism for these types of cancer, so we believe this is how we will deliver the greatest impact from our project. From an imaging perspective, analysis of each of these different tissue types presents unique measurement challenges, so developing new methods for each tissue is important to ensure they can be applied to as many types of cancer as possible in the future.
What imaging techniques will you be using?
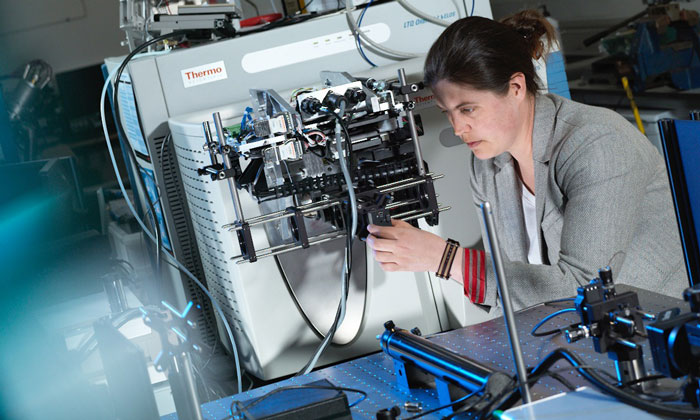
We will utilise a range of mass spectrometry imaging (MSI) techniques (MALDI, desorption electrospray ionisation mass spectrometry (DESI MS), secondary ion mass spectrometry (SIMS), inductively coupled plasma mass spectrometry (ICP-MS) and rapid evaporative ionisation mass spectrometry (REIMS)), as well as magnetic resonance spectroscopy and imaging. This will allow us to build a comprehensive, spatially resolved picture of the metabolism of solid tumours from subcellular to anatomical levels using in-vivo, ex-vivo and in-vitro analyses.
 How will you ensure patient data security and protection? Where will the data be stored?
How will you ensure patient data security and protection? Where will the data be stored?
All patient samples used in the project will be anonymised. During the early stages of the project, we’ll finalise our plans for the long-term storage of the data we generate at a later date, and we are currently exploring a whole range of possibilities.
How will you use the information gathered to better diagnose and treat cancer patients?
The aim is to create a database containing all of the gathered data, which will be freely available to researchers around the globe, for the purpose of creating a standardised way for scientists and doctors to use these new techniques in their work and in the clinic. These data could be used to identify new drug targets and will provide a better understanding of molecular changes in cancer, which may be useful for new diagnostics. We hope that this will lead to the development of better, faster ways to diagnose and treat cancer and help more people survive for longer.

What would be your advice to young people in our industry looking for funding?
I would advise people to consider both large and small funding opportunities and to not confine themselves to discipline specific calls or funding sources. And keep applying!
Related topics
Big Data, Funding, Imaging, Informatics, Magnetic resonance images (MRI), Mass Spectrometry
Related conditions
Bowel cancer, Breast cancer, Pancreatic cancer
Related organisations
Cancer Research UK, NPL
Related people
Dr Josephine Bunch


 How will you ensure patient data security and protection? Where will the data be stored?
How will you ensure patient data security and protection? Where will the data be stored?