Potential therapeutic protein inhibitors for Chagas disease
Posted: 26 September 2017 | Dr Zara Kassam (Drug Target Review) | No comments yet
Researchers have used a multi-modal integrated approach to develop potential new anti-Chagas therapies…

Researchers have used a multi-modal integrated approach to develop potential new anti-Chagas therapies by combining the principles of structure-based drug design, where therapy is designed with the knowledge of the target’s three-dimensional (3D) structure, with in vitro testing methods, and X-ray crystallography.
This approach narrows the range of potential drug candidates more efficiently.
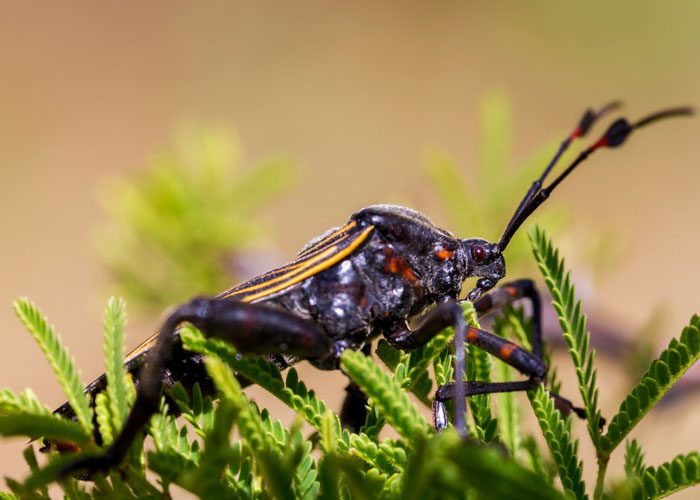
Chagas disease is a potentially life-threatening illness caused by the parasite, Trypanosoma cruzi, and is transmitted to humans through triatomine blood-sucking bugs.
Through virtual screening researchers at Nagasaki University and selected their target protein, T. cruzi spermidine synthase, based on specific structural features and properties indicating its importance for survival in another Trypanosoma species.
If the protein is required for survival of a species, inhibiting that protein could be a potential mechanism of action for a drug with activity against the parasite that causes Chagas disease.
The researchers focused on Spermidine synthase (SpdSyn) as the target protein. Following selection, potential drug candidate inhibitors were identified through a screening search known as docking simulation–a structure-based drug design approach using 3D simulations to computationally match drug compounds to SpdSyn.
They successfully identified four drug-like compounds that were virtual “matches” then evaluated their inhibition activity in vitro and compared the results with those of a positive control.
To further test potential activity and binding, they employed X-ray crystallography to confirm these four compounds in complex with the protein structure. Through interaction analyses for each compound, the researchers found that all four compounds interacted with the proposed target binding sites through the same amino acid, Asp171.
Additionally, molecular simulation suggested additional interacting sites for each compound that was not predicted by docking simulation.
Dr Masakazu Sekijima of Advanced Computational Drug Discovery Unit, at Tokyo Tech, and team believe that their study’s findings are indicative of the promise that docking simulation holds for the identification of potential drug-like inhibitors of the target protein and therapies for Chagas disease.
They hope to demonstrate the general applicability of their approach, opening doors to the discovery of treatments for other diseases.
Related topics
Drug Discovery, Drug Discovery Processes, Drug Targets, Proteomics, X-ray Crystallography
Related conditions
Chagas disease
Related organisations
Kiyoshi Kita of School of Tropical Medicine, Nagasaki University, Tokyo Tech
Related people
Dr Masakazu Sekijima