X-rays reveal structure of SARS-CoV-2 protease at room temperature
Posted: 29 June 2020 | Victoria Rees (Drug Target Review) | 1 comment
Researchers grew large crystals and used an X-ray machine with a less intense beam to elucidate the structure of the SARS-CoV-2 main protease at room temperature.
A team of researchers has performed the first room-temperature X-ray measurements of the SARS-CoV-2 main protease – the enzyme that enables the COVID-19 virus to reproduce. According to the team, these measurements mark an important first step in their ultimate goal of building a comprehensive three-dimensional (3D) model of the enzymatic protein.
The model will be used to advance supercomputing simulations aimed at finding drug inhibitors to block the SARS-CoV-2 replication mechanism and help end the COVID-19 pandemic.
The study was conducted by the US Department of Energy’s Oak Ridge (ORNL) and Argonne national laboratories.
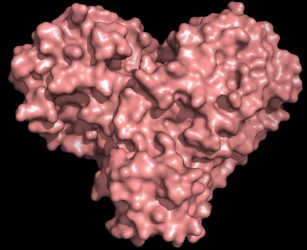
The protease protein is both shaped like a heart and functions as one, allowing the virus to replicate and spread. Inhibiting the protease would block virus reproduction [credit: Andrey Kovalevsky/ORNL].
“The protease is indispensable for the virus life-cycle. The protein is shaped like a valentine’s heart, but it really is the heart of the virus that allows it to replicate and spread. If you inhibit the protease and stop the heart, the virus cannot produce the proteins that are essential for its replication. That is why the protease is considered such an important drug target,” said ORNL’s Andrey Kovalevsky, the corresponding author of the study. While the structure is known from cryogenically preserved crystals, he added: “this is the first time the structure of this enzyme has been measured at room temperature, which is significant because it is near the physiological temperature where the cells operate.”
Due to the intensity of X-ray beams at most large-scale synchrotron facilities, biological samples typically must be cryogenically frozen to around 100K, or approximately -280°F, to withstand the radiation long enough for data to be collected.
To extend the lifetime of the SARS-CoV-2 crystallised protein samples and measure them at room temperature, the researchers grew crystals larger than required for synchrotron cryo-studies and used an in-house X-ray machine that features a less intense beam.

SARS-CoV-2 protease crystals, grown in ORNL’s Protein Crystallization and Characterization laboratory and pictured in microscopic view, will be used in X-ray scattering experiments [credit: Daniel Kneller/ORNL].
“Growing protein crystals and collecting data is a tedious and time-consuming process. In the time it typically takes to prepare and ship the sample to a synchrotron, we were able to grow the crystals, take the measurements and begin analysing the data,” said ORNL’s Daniel Kneller, the study’s first author.
The researchers say the SARS-CoV-2 protease enzyme consists of chains of amino acids with a repeating pattern of nitrogen-carbon-carbon atoms that form the backbone of the protein. Side groups of the amino acid building blocks, or ‘residues’, extend from each of the central backbone carbon atoms. The enzyme is folded into a specific 3D shape, creating special pockets where a drug molecule could attach.
The study revealed significant structural disparities between the orientations of the backbone and some of the residues in the room-temperature and cryogenic samples. The research suggests that freezing the crystals may introduce structural artefacts that could result in a less accurate understanding of the protease structure.
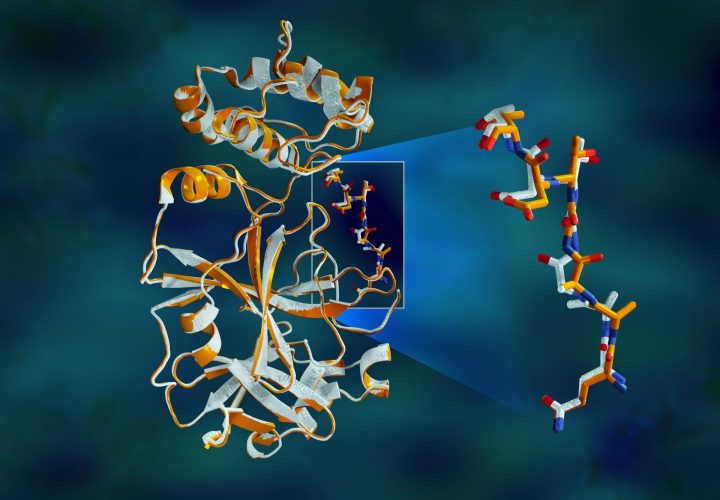
Overlapping X-ray data of the SARS-CoV-2 main protease shows structural differences between the protein at room temperature (orange) and the cryogenically frozen structure (white) [credit: Jill Hemman/ORNL].
The team’s results are being shared with researchers, led by ORNL-University of Tennessee Governor’s Chair Jeremy Smith, who are conducting drug docking simulations using Summit at ORNL – the fastest supercomputer in the US.
“What researchers are doing on Summit is taking known drug compounds and trying to computationally bind them to the main protease for drug repurposing, as well as looking for new leads into other potential drug candidates,” said ORNL corresponding author Leighton Coates. “Our room temperature data is being used to build a more accurate model for those simulations and improve drug design activities.”
The researchers’ next step in completing the 3D model of the SARS-CoV-2 main protease is to use neutron scattering at ORNL’s High Flux Isotope Reactor and the Spallation Neutron Source. Neutrons are essential in locating the hydrogen atoms, which play a critical role in many of the catalytic functions and drug design efforts.
The study can be found in Nature Communications.
Related topics
Disease Research, Drug Targets, Imaging, Research & Development, Structural Biology, Targets, X-ray Crystallography
Related conditions
Coronavirus, Covid-19
Related organisations
Argonne National Laboratory, Oak Ridge National Laboratory (ORNL), University of Tennessee
Related people
Andrey Kovalevsky, Daniel Kneller, Jeremy Smith, Leighton Coates




Thanks to the writer for sharing this blog post it’s very helpful to me