New fluorescence imaging method enables live RNA visualisation
Posted: 26 February 2021 | Victoria Rees (Drug Target Review) | No comments yet
A fluorescence imaging technique has allowed scientists to observe RNA in real time using single-molecule localisation microscopy.

To achieve a more detailed understanding of the precise function that RNA performs, researchers at Heidelberg University and the Karlsruhe Institute of Technology (KIT), both Germany, have devised a new fluorescence imaging method which provides live-cell RNA visualisation with unprecedented resolution.
The method is based on a novel molecular marker called Rhodamine-Binding Aptamer for Super-Resolution Imaging Techniques (RhoBAST). This RNA-based fluorescence marker is used in combination with the dye rhodamine. Due to their distinctive properties, the marker and dye interact in a very specific way, which makes individual RNA molecules glow. They can then be made visible using single-molecule localisation microscopy (SMLM), a super-resolution imaging technique. Due to a lack of suitable fluorescence markers, direct observation of RNA via optical fluorescence microscopy has been severely limited to date.
The marker created for RhoBAST is genetically encodable, which means that it can be fused to the gene of any RNA produced by a cell. RhoBAST itself is non-fluorescent, but lights up a cell-permeable rhodamine dye by binding to it in a very specific way.
“This leads to a dramatic increase in fluorescence achieved by the RhoBAST-dye complex, which is a key requirement for obtaining excellent fluorescence images,” explained Dr Murat Sunbul, one of the researchers from the IPMB. “However, for super-resolution RNA imaging the marker needs additional properties.”
The researchers discovered that each rhodamine dye molecule remains bound to RhoBAST for approximately one second only before becoming detached again. Within seconds, this procedure repeats itself with a new dye molecule.
“It is quite rare to find strong interactions – as between RhoBAST and rhodamine – combined with exceptionally fast exchange kinetics,” said Professor Gerd Ulrich Nienhaus, another of the researchers from the APH. Since rhodamine only lights up after binding to RhoBAST, the constant string of newly emerging interactions between marker and dye results in ‘blinking’.
“This ‘on-off switching’ is exactly what we need for SMLM imaging,” continued Nienhaus.
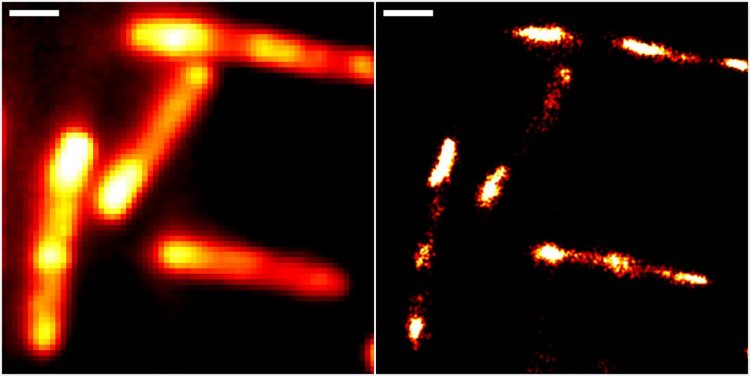

Conventional epifluorescence (left) and super-resolved localisation microscopy images of gut bacteria (Escherichia coli), using the new RhoBAST-dye marker complex for fluorescence labelling. Scale bar: 1 μm [credit: Heidelberg University/Karlsruhe Institute of Technology].
As the fluorescence images are collected under laser light irradiation, this destroys the dye molecules over time. The fast dye exchange ensures that photobleached dyes are replaced by fresh ones. This means that individual RNA molecules can be observed for longer periods of time, which can greatly improve image resolution.
The researchers were able to demonstrate the properties of RhoBAST as an RNA marker by visualising RNA structures inside Escherichia coli and cultured human cells with excellent localisation precision.
“We can reveal details of previously invisible subcellular structures and molecular interactions involving RNA using super-resolution fluorescence microscopy. This will enable a fundamentally new understanding of biological processes,” said Professor Andres Jaeschke, a scientist at the IPMB.
The study was published in Nature Biotechnology.
Related topics
Imaging, Microscopy, RNAs
Related organisations
Heidelberg University, Karlsruhe Institute of Technology (KIT)
Related people
Dr Murat Sunbul, Professor Andres Jaeschke, Professor Gerd Ulrich Nienhaus