HAT proteins could unveil new cancer and Alzheimer’s treatments
Posted: 30 November 2021 | Anna Begley (Drug Target Review) | No comments yet
The new study used cryo-electron microscopy to visualise the structure of a HAT protein and uncover compounds for drug development.

A team at at the Spanish National Cancer Research Center (CNIO) and the Institute for Research in Biomedicine (IRB Barcelona), both Spain, have visualised the structure of a member of the HAT family at atomic resolution and determine the pocket where these proteins bind to amino acids, as well as the details of the mechanism by which this recognition occurs. Their findings could lead to compounds that could act on specific regions of these proteins to treat illnesses such as cancer and neurodegenerative diseases like Alzheimer’s disease.
NEWS: Existing cancer drugs could become new Alzheimer’s treatments
Although HAT proteins are almost identical in structure, some transport certain amino acids and not others, thereby conferring each member of the family specific functions, such as the participation in cell growth; a role in diseases such as cancer; neuronal functions; and the transport of toxic substances and involvement in addiction to substances such as cocaine.
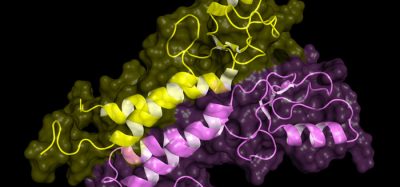
To understand this specificity of function, the team set out to study the three-dimensional (3D) structure of this family of proteins. “Classical techniques used to determine the structure of proteins, such as those using X-rays, have had limited success with proteins that are embedded in biological membranes, and so many questions have remained unresolved,” explained Dr Oscar Llorca.
“The combination of structural resolution by cryo-electron microscopy with molecular dynamics calculations and functional studies provides an experimental platform with a lot of potential that allows us to unravel the function of amino acid transporters. In this case, we have applied this technology to identify the molecular mechanisms that lead these proteins to transport some amino acids but not others,” added researcher Professor Manuel Palacín.
Thanks to their findings, published in PNAS, the researchers have been able to visualise the structure of a member of the HAT family at atomic resolution and determine the pocket where these proteins bind to amino acids, as well as the details of the mechanism by which this recognition occurs.
The atomic details reveal that only a few residues of these proteins determine the amino acids to which they bind and therefore their specific functions. In addition, the study demonstrates how the substitutions of some residues for others in these positions in the different members of the family are responsible for modifying the specificity of recognition and transport of some amino acids and not others.
The team stated that the results of this research will now allow efforts to be directed towards compounds that could act on specific regions of these proteins, and to manage the disorders in which they participate, such as cancer and neurodegenerative diseases like Alzheimer’s disease.
Related topics
Analytical Techniques, Drug Development, Drug Leads, Microscopy, Protein, Protein Expression, Small Molecules
Related conditions
Alzheimer's disease (AD), Cancer
Related organisations
Institute for Research in Biomedicine (IRB Barcelona), Spanish National Cancer Research Center (CNIO)
Related people
Manuel Palacín, Oscar Llorca