Machine learning calculates affinities of drug candidates and targets
Posted: 16 March 2021 | Victoria Rees (Drug Target Review) | No comments yet
A novel machine learning method called DeepBAR could accelerate drug discovery and protein engineering, researchers say.

A new technology combining chemistry and machine learning could aid researchers during the drug discovery and screening process, according to scientists at MIT, US.
The new technique, called DeepBAR, quickly calculates the binding affinities between drug candidates and their targets. The approach yields precise calculations in a fraction of the time compared to previous methods. The researchers say DeepBAR could one day quicken the pace of drug discovery and protein engineering.
“Our method is orders of magnitude faster than before, meaning we can have drug discovery that is both efficient and reliable,” said Professor Bin Zhang, co-author of the study’s paper. The affinity between a drug molecule and a target protein is measured by a quantity called the binding free energy – the smaller the number, the better the bind. “A lower binding free energy means the drug can better compete against other molecules, meaning it can more effectively disrupt the protein’s normal function.”
Calculating the binding free energy of a drug candidate provides an indicator of a drug’s potential effectiveness. However, it is a difficult quantity to discover. Methods for computing binding free energy fall into two broad categories:
- One calculates the quantity exactly, eating up significant time and computer resources
- The second is less computationally expensive, but it yields only an approximation of the binding free energy.
The researchers devised an approach to get the best of both worlds. DeepBAR computes binding free energy exactly, but requires just a fraction of the calculations demanded by previous methods.
The “BAR” in DeepBAR stands for “Bennett acceptance ratio,” a decades-old algorithm used in exact calculations of binding free energy. Using the Bennet acceptance ratio typically requires a knowledge of two “endpoint” states, eg, a drug molecule bound to a protein and a drug molecule completely dissociated from a protein, plus knowledge of many intermediate states, eg, varying levels of partial binding, all of which slow down calculation speed.
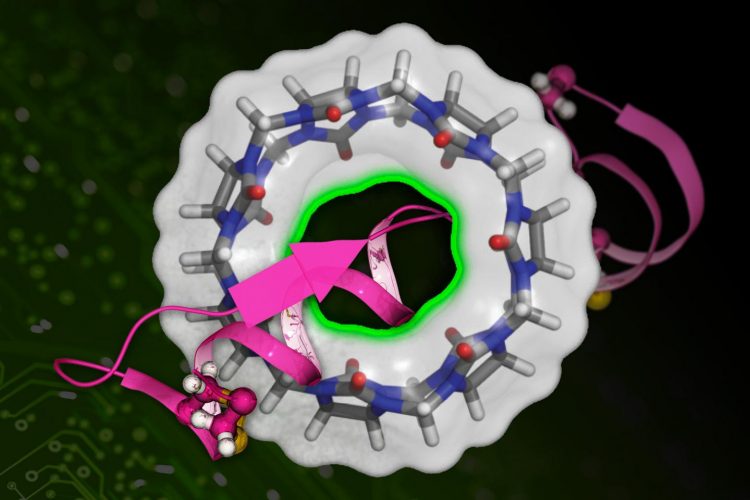
MIT researchers have developed a machine learning-based technique to more quickly calculate the binding affinity of a drug molecule (represented in pink) with a target protein (the circular structure) [credit: MIT News, Xinqiang Ding and Bin Zhang].
DeepBAR reduces in-between states by deploying the Bennett acceptance ratio in machine learning frameworks called deep generative models.
“These models create a reference state for each endpoint, the bound state and the unbound state,” said Zhang. These two reference states are similar enough that the Bennett acceptance ratio can be used directly, without all the costly intermediate steps.
“It is basically the same model that people use to do computer image synthesis,” says Zhang. “We are sort of treating each molecular structure as an image, which the model can learn. So, this project is building on the effort of the machine learning community.”
“These models were originally developed for two-dimensional (2D) images,” said lead author of the study Xinqiang Ding. “But here we have proteins and molecules – it is really a three-dimensional (3D) structure. So, adapting those methods in our case was the biggest technical challenge we had to overcome.”
In tests using small protein-like molecules, DeepBAR calculated binding free energy nearly 50 times faster than previous methods. The researchers add that, in addition to drug screening, DeepBAR could aid protein design and engineering, since the method could be used to model interactions between multiple proteins.
In the future, the researchers plan to improve DeepBAR’s ability to run calculations for large proteins, a task made feasible by recent advances in computer science.
“This research is an example of combining traditional computational chemistry methods, developed over decades, with the latest developments in machine learning,” said Ding. “So, we achieved something that would have been impossible before now.”
The research is published in Journal of Physical Chemistry Letters.
Related topics
Artificial Intelligence, Drug Discovery, Informatics, Molecular Targets, Screening
Related organisations
MIT
Related people
Professor Bin Zhang, Xinqiang Ding