A new computational tool, DeepTarget, is demonstrating context-specific targets and repurposing opportunities, showing that what may be a side effect in one patient could serve as a treatment in another.
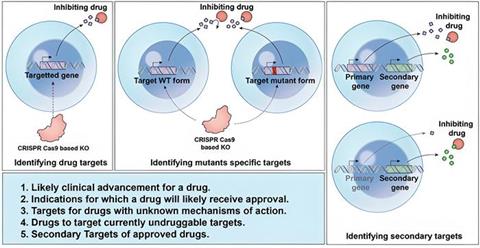
A new study from scientists at Sanford Burnham Prebys suggests that what is considered a side effect in one patient could provide a treatment in another, if researchers broaden their perspective on small molecule drug targets.
Small molecules: more than one target
Looking at the bigger picture shows that small molecules can act on different targets and produce varying effects depending on the disease and cell type, offering opportunities to repurpose drugs for more patients.
“The kinds of small molecules representing many of our medicines are rarely found in nature, so they haven’t evolved to carry out a specific task,” said Dr Sanju Sinha, an assistant professor in the Cancer Metabolism and Microenvironment Program at Sanford Burnham Prebys Medical Discovery Institute. “Sometimes the field looks at these drugs with tunnel vision in terms of them having a single target along with some side effects labelled as ‘off-target effects.’
DeepTarget: a new computational tool
Dr Sinha began exploring the flexibility of small molecule drugs during his training at the National Cancer Institute. He developed a computational tool called DeepTarget, which predicts drug targets not based on chemical structures, but using large-scale genetic and drug screening data from cancer cells. The dataset analysed by Sinha and his collaborators included information on 1,450 drugs across 371 cancer cell lines, sourced from the Dependency Map (DepMap) Consortium.
Being able to predict these secondary targets is important because many FDA-approved drugs and new drugs in clinical development have them.
In tests comparing DeepTarget’s predictions of primary cancer drug targets with existing drug-target data, the tool outperformed current state-of-the-art methods – including RoseTTAFold All-Atom and Chai-1 – in seven out of eight scenarios. It also successfully predicted whether drugs preferentially affected typical, non-mutated target proteins or their mutant forms, as well as identifying secondary targets.
“Being able to predict these secondary targets is important because many FDA-approved drugs and new drugs in clinical development have them,” said Sinha, lead author of the study. “If we can see them more as features rather than bugs, we can take advantage of these targets to improve drug repurposing.”
Case study: Ibrutinib and lung cancer
To validate DeepTarget’s predictions, the team conducted two experimental case studies, including one involving Ibrutinib, an FDA-approved treatment for blood cancer. Previous clinical research had indicated that Ibrutinib could also treat lung cancer, despite its primary target, Bruton’s tyrosine kinase (BTK), not being present in lung tumours.
Cells carrying the mutant EGFR were more sensitive to Ibrutinib, confirming EGFR as a relevant target.
In collaboration with co-corresponding author Dr Ani Deshpande, a professor in the Cancer Genome and Epigenetics Program at Sanford Burnham Prebys, the researchers tested DeepTarget’s whether Ibrutinib affected lung cancer cells by targeting a secondary protein, epidermal growth factor receptor (EGFR).
“In consulting DeepTarget, if we only focused on blood tumours, then BTK was the primary target,” said Sinha. “If we changed our focus to solid tumours, then a mutant, oncogenic form of EGFR became the primary target, so this was a clear example of a context-specific target.”
Cells carrying the mutant EGFR were more sensitive to Ibrutinib, confirming EGFR as a relevant target.

DeepTarget’s predictions are based on the principle that removing a gene encoding the protein target of a given drug through CRISPR-Cas9 gene editing can mimic the inhibitory effects of that drug. The tool was built by leveraging large-scale genetic and drug screening experiments with comprehensive data for 1450 drugs across 371 cancer cell lines. Credit: Sanju Sinha, Sanford Burnham Prebys.[/caption]
Implications for drug development
“We believe that the tool’s superior performance in real-world scenarios is due to it more closely mirroring real-world drug mechanisms, where cellular context and pathway-level effects often play crucial roles beyond direct binding interactions,” said Sinha. “It also underscores DeepTarget’s potential to accelerate drug development and repurposing efforts as a complementary approach alongside structural methods focused on chemical binding.”
We believe that the tool’s superior performance in real-world scenarios is due to it more closely mirroring real-world drug mechanisms.
The study highlights a shift in how researchers approach drug development, emphasising the importance of understanding cellular context and secondary targets. By embracing these ideas, tools like DeepTarget could accelerate the discovery of new treatments. Sinha now hopes to leverage these insights to create new small molecule drugs.
“Improving treatment options for cancer and for related and even more complex conditions like aging will depend on us improving both our ways to understand the biology, as well as ways to modulate it with therapies,” he said.
Topics
- Artificial Intelligence (AI)
- Cancer
- Cancer
- Computational Techniques
- Dr Sanju Sinha (assistant professor in the Cancer Metabolism and Microenvironment Program at Sanford Burnham Prebys Medical Discovery Institute)
- Drug Discovery
- Drug Discovery Processes
- Drug Repurposing
- Drug Targets
- High-Throughput Screening (HTS)
- Molecular Biology
- Molecular Targets
- Sanford Burnham Prebys
- Small Molecules
- Tools and techniques
- Translational Science