For decades, drugging the ‘undruggable’ was thought to require luck rather than logic. Today, AI is transforming serendipity into strategy by enabling rational, data-driven approaches to previously inaccessible targets.

Many targets have been considered undruggable by traditional means. Until recently, many often viewed these targets with cautious excitement, believing that making them druggable required serendipity, not rational design. The prevailing dogma was that these targets couldn’t possibly be selective. However, by combining artificial intelligence (AI) algorithms, diverse libraries and the ability to ‘read’ the library, alongside omics technologies, the target space can now be expanded beyond these historical limitations.
Serendipity: from chance to breakthroughs
Drug discovery has long been intertwined with serendipity, ie, scientific discoveries that started from accidental events. Classic examples include penicillin and cisplatin, which were both accidentally discovered by Alexander Fleming and Barnett Rosenberg, respectively. Serendipity happens in different ways as accidental events can differently impact on scientific research. However, there is more to serendipity than pure chance, even in the case of ‘untargeted research’ and no scientific discovery has ever been achieved by pure luck. Indeed, by virtue of intellectual preparation, experience, curiosity, sagacity and imagination, scientists were able to profit from casual occurrences, often taking advantage of unexpected results that paved the way to breakthroughs. Now, AI has the power to fundamentally revolutionise this process by replacing much of both the accidental element and the human intellectual effort with advanced computational models.
Computational rationality and the risk of guided innovation
Computational rationality describes an approach that leverages mathematical and algorithmic models to make optimal decisions, accounting for real-world constraints in both data and computation. In drug discovery, such an approach enables AI algorithms to optimise decisions using existing pharmacological data, significantly enhancing efficiency and speed. Moreover, AI facilitates the development of highly personalised therapies: advanced analysis of genomic and clinical data enables drugs tailored to specific genetic subgroups, improving treatment efficacy and reducing side effects.
Heavy reliance on existing data may limit the exploration of novel and unpredictable compounds, which have historically driven medical breakthroughs.
However, these advantages come with trade-offs. Heavy reliance on existing data may limit the exploration of novel and unpredictable compounds, which have historically driven medical breakthroughs. AI systems risk overlooking unconventional molecules or drugs with atypical mechanisms of action, potentially leading the pharmaceutical industry to converge on similar, predictable solutions and reducing therapeutic diversity.
Additionally, the predictability of algorithmic decision-making may suppress the role of serendipity – an element that has often catalysed revolutionary discoveries. The influence of randomness in drug development is far from negligible; many widely used medications are the result of unexpected findings and fortunate accidents.
High-quality dataset
The effectiveness of AI models hinges directly on the availability of robust, high-quality data. While AI has made significant strides in designing small molecules that effectively target specific proteins, this success is largely attributable to the existence of one meticulously curated dataset: the Protein Data Bank (PDB). For decades, spanning generations of scientists since the 1950s, the PDB has served as the central archive for protein structures. This dedication to accuracy and completeness has yielded a database of exceptional depth and utility.
The PDB’s value lies not just in its sheer volume of data, but in its rigorous curation. Every entry undergoes careful validation, ensuring the reliability of the information it contains. This high standard of quality is crucial for training AI models, as it minimises the introduction of errors and biases that could compromise the accuracy of predictions. Consequently, AI-driven drug discovery efforts leveraging the PDB have achieved notable breakthroughs, particularly in identifying potential drug candidates that interact with specific protein targets. This success story underscores a foundational principle: AI performance in terms of accountability, tabular and semantic explainability, reliability, robustness, generalisation and accuracy, is directly proportional to the quality and quantity of the data it is trained on.
(Un)druggable targets
A significant portion of disease-relevant proteins remains ‘undruggable’ by traditional therapeutic approaches, posing a major challenge in drug discovery. AI promises more than just the optimisation of known chemical and biological spaces; it is meant to actively explore uncharted territories. Recent advances in AI algorithms, such as AlphaFold2 and RoseTTAFold, promise to transform several previously undruggable targets into druggable ones by overcoming long-standing barriers in drug design. AlphaFold 3, co-developed by Isomorphic Labs and Google DeepMind, represents a recent step change in protein structure prediction. This new era blurs the line between druggable and undruggable targets, effectively expanding the druggable proteome and opening new therapeutic possibilities for diseases once considered untreatable, including cancer, neurological disorders and metabolic diseases.
Beyond the binding site
Drugging the undruggable lies in AI’s ability to create entirely new strategies beyond traditional occupancy-based inhibition. For decades, approximately 85 percent of disease-causing proteins were considered undruggable because they lack the well-defined binding pockets required for small-molecule drugs. A key part of AI’s promise is its ability to uncover cryptic pockets – binding sites that are absent in a protein’s natural state but appear transiently or when a ligand binds. By revealing these previously ‘invisible’ sites, AI is creating new entry points for drug design on targets once considered completely intractable. In addition to finding new sites, AI is dismantling this barrier by enabling the rational design of novel therapeutic modalities that function through different mechanisms.
Drugging the undruggable lies in AI’s ability to create entirely new strategies beyond traditional occupancy-based inhibition.
Two prominent examples of these new modalities are targeted protein degraders and molecular glues. By moving beyond direct enzymatic inhibition and leveraging AI to rationally design molecules that co-opt cellular machinery, researchers are systematically expanding the druggable proteome. However, despite its immense promise, the future success of AI in drugging the undruggable depends on overcoming key challenges. The primary obstacle is the scarcity of high-quality, large-scale data needed to train and validate these sophisticated models – even though progress has been made for some modalities. The path forwards requires a coordinated effort to build specialised databases and, crucially, to establish a feedback loop where Al predictions are continuously tested through experiments, with the results used to refine and improve the next generation of Al models. This need for systemic, coordinated effort is also highlighted by the insights synthesised from the Cancer Moonshot workshop, which specifically calls for a federated database for undruggable targets and sustained, coordinated investment. The integration of cutting-edge computational tools with experimental validation holds the ultimate promise of expanding the druggable universe.
A new method for understanding protein structure and surface involves geometric deep learning. This goes beyond traditional 2D image analysis, enabling a 3D framework that captures the curvature, shape, polarity and hydrophobicity of proteins. This multidimensional model enables more accurate representation and prediction of how small molecules will interact with a target.
AI in computational drug discovery
AI also supports the discovery and optimisation of small-molecule inhibitors targeting previously undruggable pockets. Computational methods currently being enhanced by AI include molecular docking, virtual screening and simulated molecular dynamics. These methods virtually evaluate molecular binding possibilities and simulate the complex, dynamic behaviour of molecules in silico. Generative chemistry methods using AI can propose novel compounds that may not be immediately apparent to human chemists, though these still require experimental validation. For example, a multimodal transformer model can predict molecular structures using input from spectroscopic data such as infrared (IR) or nuclear magnetic resonance (NMR) spectra. Additionally, conditional variational autoencoder (CVAE) generative models are used for generating novel molecules. Together, these AI-driven computational advances are rapidly transforming the landscape of drug discovery, revealing new opportunities to effectively target proteins once deemed beyond reach.
Unexpected insights from complex models: emergent properties
Sometimes, generative AI models may seem to have learnt more than they were taught, as they exhibit behaviours, perform tasks, draw conclusions and make predictions surprisingly and unexpectedly. In particular, large language models (LLMs), characterised by a complex and multilayered architecture, are acknowledged to display emergent abilities in analogy with natural complex systems, whose emergent, distinctive properties are neither reducible to, nor predictable from, the properties of the system’s components. Several abilities of LLMs have been shown to emerge suddenly and unpredictably as the number of parameters and the size of training datasets exceed certain thresholds. This is consistent with the view that emergence occurs when quantitative changes in a system lead to qualitative changes in behaviour.
Sometimes, generative AI models may seem to have learnt more than they were taught, as they exhibit behaviours, perform tasks, draw conclusions and make predictions surprisingly and unexpectedly.
Similarly to what happens in natural systems, inherent complexity hinders traditional mechanist explanations as well as specific predictions on further abilities or behaviours that could arise in the future, thus also making it difficult to anticipate the consequences of a model’s development and deployment. Yet many of the abilities recognised as emergent do not display spontaneously in generic testing; rather they are triggered as the model is engaged in specific operational contexts. Although not intentionally constructed, emergent abilities can be intentionally induced, for instance by means of post-training techniques like prompt and fine tuning strategies and to a certain extent canalised in the wider framework of cost-effectively enhancing models’ performances without further up-scaling or re-training. Surprising, unexpected outcomes may therefore derive from the adopted methodology addressed to unlock latent yet triggerable capacities of the model for the sake of maximising its potential.
Generalisation, emergence and the future pipeline
Structural prediction serves as a powerful example of how a comprehensive, carefully curated database can accelerate scientific progress, unlocking AI’s potential in critical fields like drug discovery. To fully realise this transformative potential across diverse scientific disciplines, the development and maintenance of similar high-quality datasets must be prioritised. Aspects to consider:
- Will the industry be able to do this in a generalisable way? In other words, can this be achieved regardless of the type of target or modality?
- Will the efficient design of drugs facilitate downstream multimodal analysis-based systems that are being used to identify biomarkers and relevant patient populations?
- Will the expansion of druggable space be determined by GenAI reassembling existing data into original components or by GenAI enacting emergent abilities, where the model has in fact never seen the data before?
According to recent reports, pharmaceutical companies are investing in connected, digitalised and sustainable laboratories, with the aim of accelerating time-to-market, optimising costs and improving probability of success. AI plays a key role in automating complex processes and reducing experimental errors. However, data management, the increasing complexity of processes and the difficulty of finding professionals with cross-functional skills are significant obstacles to overcome. The direction is clear: the laboratories of the future will be highly technological environments, where AI and human intervention will collaborate to radically transform the discovery of new targets.
Concluding remarks: balancing efficiency and creativity
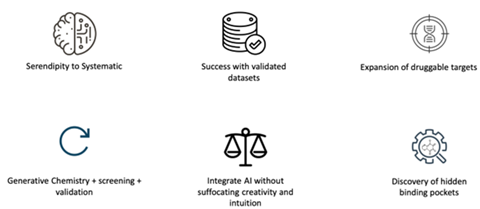
AI is making the process of pharmaceutical discovery faster and more efficient. However, to foster true innovation, these advantages should be balanced with exploring the uncharted, adopting unconventional approaches and even maintaining a certain degree of unpredictability (Figure 1).
Literature
- Akinsanya K, AlQuraishi M, Boija A, et al. 2025. Redefining druggable targets with artificial intelligence. Nat Biotechnol 43, 1416–1418 . https://doi.org/10.1038/s41587-025-02770-1
- Anderson PW. 1972. More is Different. Science 177 (4047): 393-396. https://www.science.org/doi/10.1126/science.177.4047.393
- Ban TA. 2006. The Role of Serendipity in Drug Discovery, Dialogues in Clinical Neuroscience 8 (3): 335-344. https://pubmed.ncbi.nlm.nih.gov/17117615/
- Berti L, et al. 2025. Emergent Abilities in Large Language Models: A Survey. https://arxiv.org/abs/2503.05788
- Ganguli D, et al. 2022. Predictability and Surprise in Large Generative Models. https://dl.acm.org/doi/10.1145/3531146.3533229
- Hargrave-Thomas E, et al. 2012. Serendipity in Anticancer Drug Discovery. World Journal of Clinical Oncology, 3 (1): 1-6. https://pubmed.ncbi.nlm.nih.gov/22247822/
- Steinhardt J. 2022. Future ML Systems Will Be Qualitatively Different. . https://www.alignmentforum.org/posts/pZaPhGg2hmmPwByHc/future-ml-systems-will-be-qualitatively-different
- Wei J, et al. 2022. Emergent Abilities of Large Language Models. https://arxiv.org/abs/2206.07682
- Yaqub O. 2018. Serendipity: Towards a Taxonomy and a Theory. Research Policy 47: 169-179. https://www.sciencedirect.com/science/article/pii/S0048733317301774
- Building the next-gen pharma lab DIGITALLY CONNECTED, ENVIRONMENTALLY SUSTAINABLE. CapGemini (2024) https://www.capgemini.com/wp-content/uploads/2024/02/BUILDING-THE-NEXT-GEN-PHARMA-LAB-160224-Web.pdf
- Romanelli V, Annunziata D, Cerchia C, et al. Enhancing De Novo Drug Design across Multiple Therapeutic Targets with CVAE Generative Models. ACS Omega 2024 9 (43), 43963-43976 https://pubs.acs.org/doi/10.1021/acsomega.4c08027
- Geukes Foppen RJ, Gioia V, Shreya G, et al. Methodology for Safe and Secure AI in Diabetes Management in Journal of Diabetes Science and Technology, 19, 620–627. (2025). https://doi.org/10.1177/19322968241304434
- Geukes Foppen RJ, Gioia V, Zoccoli A, Velez CN. From siloed data to breakthroughs: multimodal AI in drug discovery [Internet]. Drug Target Review. 2025. Available from: https://www.drugtargetreview.com/article/160597/from-siloed-data-to-breakthroughs-multimodal-ai-in-drug-discovery/
- Geukes Foppen RJ, Gioia V, Zoccoli A, Velez CN. The Rise of Multimodal Language Models in Drug Development. (2025) European Pharmaceutical Review Featured Article - 12 June 2025 https://www.europeanpharmaceuticalreview.com/article/256440/the-rise-of-multimodal-language-models-in-drug-development/
- 2025 the evolution of labs report. Pistoia Alliance. https://marketing.pistoiaalliance.org/hubfs/Lab%20Of%20The%20Future%20Reports/FINAL%202025%20LOTF%20Survey%20Results%20%281%29.pdf
- Shrivastav P, Singh R, Wiemer AJ. Molecular glues evolve from serendipity to rational design. Trends Pharmacol Sci. 2025 Dec 3:S0165-6147(25)00237-8. https://www.sciencedirect.com/science/article/abs/pii/S0165614725002378
- Gainza P, Bunker RD,Townson SA, Castle JC. Machine learning to predict de novo protein–protein interactions, Trends in Biotechnology, Volume 43, Issue 12, 2025, Pages 3056-3070, ISSN 0167-7799, https://doi.org/10.1016/j.tibtech.2025.04.013.
- Gershman SJ, Horvitz EJ, Tenenbaum JB. Computational Rationality: A Converging Paradigm for Intelligence in Brains, Minds, and Machines, Science 349 (pp. 273-278), 2015. https://www.science.org/doi/10.1126/science.aac6076
- Geukes Foppen RJ, Gioia V, Zoccoli A, Velez CN. Using clinical genomics and AI in drug development to elevate success. (2025) Drug Target Review, February Edition https://www.drugtargetreview.com/article/155906/clinical-genomics-ai-drug-success/
- Geukes Foppen RJ, Gioia V, Zoccoli A. Token‑Level Attribution for Transparent Biomedical AI, Biomedical Engineering and Computational Biology (2026). Vincenzo Gioia, Alessio Zoccoli, Remco Jan Geukes Foppen https://journals.sagepub.com/doi/10.1177/11795972251407864
Topics
- Analytical Techniques
- Artificial Intelligence (AI)
- Bioinformatics
- Companies
- Computational Techniques
- Drug Development
- Drug Discovery
- Drug Discovery Processes
- Drug Targets
- Genomics & Sequencing
- High-Throughput Screening (HTS)
- Imaging & Diagnostics
- Informatics
- Machine Learning (ML)
- Molecular Biology
- Molecular Modelling
- Neurological disorders
- Paolo D’Ambrosio
- Precision Medicine
- Protein Expression
- Remco Jan Geukes Foppen
- Small Molecules
- Structural Biology
- Translational Science
- Vincenzo Gioia