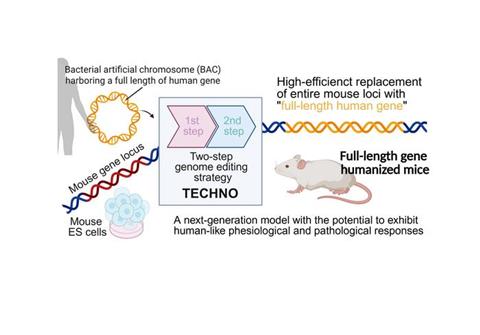
Japanese researchers have developed a new gene-editing platform, TECHNO, that allows entire human genes – including regulatory regions – to be inserted into mice, allowing for better disease modelling and drug testing.
Understanding human gene function has long been constrained by differences between species. Although mice share most protein-coding genes with humans, their regulatory landscapes often diverge, limiting how accurately mouse models can reflect human biology. A promising solution is full-length gene humanisation (FL-GH), in which entire mouse loci – including coding sequences, introns, untranslated regions and regulatory elements – are replaced with their human counterparts. However, existing technologies have struggled to insert very large genomic fragments efficiently, slowing the development of physiologically relevant humanised models.
A two-step solution: TECHNO
To tackle these challenges, a new study led by Associate Professors Dr Manabu Ozawa and Dr Jumpei Taguchi at The Institute of Medical Science, The University of Tokyo, Japan, has developed a streamlined two-step strategy for FL-GH. Their study introduces TECHNO (Two-step ES Cell-based HumaNisatiOn), a method that combines CRISPR/Cas9-assisted genome editing with bacterial artificial chromosome (BAC)-based delivery of large human genomic regions.
“This framework offers a practical and scalable solution for replacing entire mouse loci with their human counterparts,” says Dr Ozawa. “Our results demonstrate a robust and broadly applicable platform for generating FL-GH mouse models.”
How TECHNO works
The TECHNO workflow unfolds in two coordinated steps. First, the target mouse locus is excised using Cas9 ribonucleoproteins and replaced with short human homology arms surrounding a selection cassette, creating a precise genomic landing site. In the second step, a BAC carrying the full-length human gene and its regulatory elements is introduced into embryonic stem cells alongside a universal gRNA targeting the selection cassette. This enables homology-directed integration of genomic fragments exceeding 200 kilobase pairs (kbp).
Because the method relies on standard molecular reagents and widely available BAC libraries, it is theoretically applicable to more than 90 percent of human genes, offering a scalable approach to humanising the mouse genome.
 TECHNO: A two-step genome editing strategy that enables efficient replacement of entire mouse loci with full-length human genes to generate gene-humanised mice. Credit: Dr Manabu Ozawa from The University of Tokyo, Japan.[/caption]
TECHNO: A two-step genome editing strategy that enables efficient replacement of entire mouse loci with full-length human genes to generate gene-humanised mice. Credit: Dr Manabu Ozawa from The University of Tokyo, Japan.[/caption] Successful humanisation of key genes
Using TECHNO, the team successfully humanised several loci, including c-Kit, APOBEC3 and CYBB. Humanisation of c-Kit reproduced human-like alternative splicing and organ-specific expression, while supporting essential biological processes such as haematopoiesis and spermatogenesis.
Using TECHNO, the team successfully humanised several loci, including c-Kit, APOBEC3 and CYBB.
The replacement of the APOBEC3 locus demonstrated the scalability of the platform, integrating over 200 kbp of human DNA spanning seven genes and generating expression patterns that mirrored those observed in humans. Meanwhile, the humanised CYBB allele was used to introduce disease-associated mutations to model chronic granulomatous disease. The resulting mice exhibited impaired reactive oxygen species production, faithfully recapitulating the molecular phenotype seen in patients.
Implications for disease modelling and drug development
In the near term, TECHNO is expected to accelerate the development of precise, human-relevant animal models for evaluating therapeutic targets, validating disease-associated variants and identifying ineffective drug candidates earlier in research pipelines. Over the longer term, scalable FL-GH could transform biomedical research by enabling models that more accurately mimic human gene regulation and disease mechanisms.
By enabling high-efficiency integration of genomic fragments exceeding 200 kbp while preserving complex regulatory behaviour in vivo, the TECHNO platform advances the development of next-generation humanised mouse models.
“Overall, these results demonstrate that our method enables not only FL-GH of individual loci but also precise modelling of human genetic diseases in vivo by introducing disease-associated mutations into humanised alleles,” Dr Ozawa adds.
By enabling high-efficiency integration of genomic fragments exceeding 200 kbp while preserving complex regulatory behaviour in vivo, the TECHNO platform advances the development of next-generation humanised mouse models. Its versatility, robustness and reliance on standard laboratory tools make it a key technology for functional genomics and disease modelling.
Topics
- Artificial Intelligence & Computational Tools
- Cancer
- CRISPR
- Disease Research
- DNA
- Dr Jumpei Taguchi (Associate Professor at The Institute of Medical Science The University of Tokyo)
- Dr Manabu Ozawa (Associate Professor at The Institute of Medical Science The University of Tokyo)
- Drug Discovery Processes
- Drug Targets
- Genome Editing
- Genomics & Sequencing
- In Vivo Testing
- Molecular Biology
- Neurological disorders
- Technology
- The Institute of Medical Science The University of Tokyo
- Translational Science