Researchers at the University of Texas at Austin have demonstrated that targeting a bacterial protein-folding pathway can disable antimicrobial resistance and cross-protection, restoring antibiotic susceptibility in preclinical models of cystic fibrosis infection.
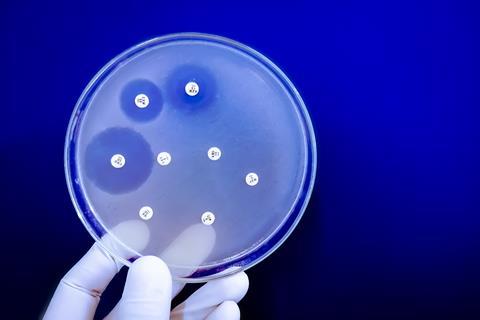
Targeting a hidden weakness in resistant bacteria
Researchers at the University of Texas at Austin have identified a new way to disable antimicrobial resistance in bacteria linked to cystic fibrosis, with potential to improve treatment of persistent infections.
The study, published in the journal eLife, describes a previously underexplored mechanism that can make antibiotic-resistant bacteria vulnerable again. By targeting a protein-folding system required for resistance, the researchers were able to restore antibiotic activity in preclinical models.
Breaking down cross-protection
Antimicrobial resistance remains one of the most pressing global health challenges, with some pathogens now resistant to nearly all available treatments. In many infections, particularly those associated with cystic fibrosis, multiple bacterial species coexist and interact in ways that further reduce the effectiveness of antibiotics.
One such interaction is known as cross-protection, where resistant bacteria degrade antibiotics in their surroundings, allowing more sensitive microbes to survive. This shared defence mechanism can make infections significantly harder to treat.
The research team, led by scientists at the University of Texas at Austin, focused on disrupting a protein-folding system required for resistance enzymes to function. These enzymes, including β-lactamases, break down widely used antibiotics such as penicillins and cephalosporins.
By removing the gene responsible for this system, the researchers found that bacteria lost their resistance and became susceptible to antibiotics. They also showed that the same effect could be achieved using chemical inhibitors, highlighting a potential pathway for drug development.
This approach is illustrated below, showing how inhibiting bacterial resistance mechanisms can restore antibiotic activity.
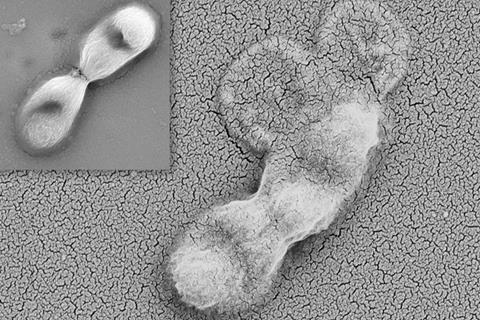
An antibiotic-resistant bacterium (Escherichia coli) is treated with an antibiotic (colistin) and a DsbA inhibitor, causing it to rupture and, eventually, die. The work highlights a new strategy that aims to eradicate multidrug-resistant pathogens. Credit: Nikol Kadeřábková
Blocking protein folding disables resistance
Experiments in mixed bacterial communities and infected wax moth larvae showed that disrupting the protein-folding system restored antibiotic sensitivity and prevented cross-protection between species.
“Certain pathogens have developed resistance to nearly all available antibiotics. In some cases, they can also shield one another, making standard treatments far less effective,” said lead author Nikol Kadeřábková, a research associate at the University of Texas at Austin. “By targeting a protein-folding system, our research shows that both resistance and cross-protection can be inactivated, allowing conventional antibiotics to regain their effectiveness.”
Certain pathogens have developed resistance to nearly all available antibiotics. In some cases, they can also shield one another, making standard treatments far less effective.
The study examined interactions between Pseudomonas aeruginosa, a common cause of cystic fibrosis lung infections and Stenotrophomonas maltophilia, a highly drug-resistant bacterium increasingly found in patients. The latter produces enzymes that can degrade antibiotics in the surrounding environment, effectively protecting neighbouring bacteria and contributing to the emergence of further resistance.
By disrupting the folding system required for these enzymes to function, the researchers were able to prevent this cross-protection.
Extending existing antibiotic activity
The findings point to a potential strategy for improving the performance of existing antibiotics by targeting shared resistance mechanisms across bacterial species.
While our work so far has been conducted in the lab, it highlights a previously untapped vulnerability in some of the most stubborn antibiotic-resistant bacteria.
“While our work so far has been conducted in the lab, it highlights a previously untapped vulnerability in some of the most stubborn antibiotic-resistant bacteria,” said Despoina Mavridou, assistant professor of molecular biosciences at the University of Texas at Austin. “By targeting the protein-folding system these pathogens rely on to build their resistance enzymes, we may be able to develop a new class of therapies that work alongside standard antibiotics, restoring their effectiveness and helping clinicians treat infections that are currently very difficult to manage.”
Similar resistance mechanisms are found across a wide range of bacterial species, suggesting broader applicability beyond cystic fibrosis. Targeting shared defence systems could provide a route to overcoming antimicrobial resistance and extending the lifespan of existing antibiotics.














No comments yet