In the wake of recent government policy aimed at actively replacing animal models in drug discovery, we consider a possible solution to the translational shortfalls of current cellular methodologies for neurodegenerative disease.
The pharmaceutical industry faces a stark reality: traditional in vitro and animal models are failing to translate into clinical success, particularly in neurodegenerative diseases. For example the attrition of Alzheimer’s therapeutics from Phase I to approval is near complete1 with high-profile failures such as Bapineuzumab, Solanezumab and Crenezumab illustrating the disconnect between preclinical promise and patient outcomes.2,3
These drugs demonstrated plaque reduction in transgenic mouse models but failed to improve cognition in humans, underscoring two critical limitations: firstly, rodent models do not capture the complexity of human neurodegeneration;4 secondly, preclinical endpoints focus on cellular biomarkers rather than functional outcomes that matter to patients, such as cognition and mobility.
Traditional in vitro and animal models are failing to translate into clinical success, particularly in neurodegenerative diseases.
Given that 2D cell cultures are widely recognised as inadequate for modelling the multicellular, chronic and age-dependent nature of neurodegeneration, the industry has increasingly turned to advanced in vitro systems, including three-dimensional organoids and perfused co-cultures or microfluidic platforms.5 Organoids recapitulate aspects of tissue architecture and cell–cell interactions that 2D systems cannot, while perfusion-enabled co-cultures further improve physiological relevance by introducing controlled flow, nutrient delivery and limited inter-tissue communication. These advances make possible experiments previously impossible in 2D, such as modelling the blood–brain barrier to assess CNS drug permeability,6,7 transcytosis and barrier disruption, or intestinal absorption8 and metabolism for early pharmacokinetic evaluation. Such systems represent a significant improvement over conventional 2D cultures, yet they remain fundamentally limited.
Organoids derived from pluripotent stem cells resemble foetal or early postnatal tissue, thus failing to capture ageing-dependent processes like mitochondrial dysfunction, epigenetic drift and chronic neuroinflammation – the very hallmarks driving neurodegenerative disease.9,10 Perfused co-cultures, while enabling multitissue interactions, fall short of replicating full cellular diversity, vascular complexity and functional neural network connectivity. Furthermore, endpoints such as cognition or motor coordination remain inaccessible, leaving a critical translational gap.
This drive towards advanced cellular methodologies is mirrored by government policy11,12. In the UK, the 2025 ‘Replacing Animals in Science’ strategy signalled a major shift from reducing or refining animal use towards actively replacing animal models wherever scientifically feasible. Significant public investment – £60 million to establish hubs for data, technology and alternative methods, plus further funding through the Medical Research Council, Innovate UK and the Wellcome Trust – is accelerating the adoption of organoids, organ-on-chip technologies, 3D bioprinted tissues and AI-driven modelling. Regulatory authorities, including the UK Centre for the Validation of Alternative Methods (UKCVAM), the European Medicines Agency (EMA) and the US Food and Drug Administration (FDA), are increasingly embedding new approach methodologies (NAMs) into drug development frameworks. These initiatives are essential for advancing human-relevant science and align with ethical imperatives to reduce mammalian testing.
Yet current NAMs efforts – focused on enhancing in vitro complexity and computational prediction – reveal blind spots. Funding and regulatory momentum gravitate towards organoids and microfluidic systems because they are direct extensions of traditional cell culture paradigms: same tissues, same endpoints – just under flow or in three dimensions. This leaves unanswered questions about organism-level responses, ageing, systemic interactions and functional outcomes. Policies aimed at replacing animals must therefore broaden their scope to include lower metazoan whole-organism models if the translational gap is to be truly addressed.
Funding and regulatory momentum gravitate towards organoids and microfluidic systems because they are direct extensions of traditional cell culture paradigms.
Lower metazoan whole-organism models – particularly invertebrates such as Caenorhabditis elegans – offer a practical solution. These organisms are small, highly fecund, ethically unregulated and amenable to genetic manipulation, allowing for scalable high-throughput screening (HTS). Critically, they enable the study of functional endpoints – paralysis, mobility, chemotaxis and survival – within the context of natural ageing and disease progression. C. elegans can model human-relevant proteinopathies such as amyloid-β, α-synuclein and polyglutamine expansions,13,14 offering a bridge between cellular readouts in advanced 3D systems and clinically meaningful functional outcomes in mammals.
Neurodegeneration is not only a disease of the brain but is influenced by systemic factors such as the gut–brain axis. Dysbiosis and altered microbial metabolites contribute to neuroinflammation, protein aggregation and functional decline in conditions including Alzheimer’s, Parkinson’s and amyotrophic lateral sclerosis (ALS).15-18 Current 3D models can partially probe these effects through supplementation with bacterial metabolites, but dynamic, live interactions between microbiota, gut epithelium and neuronal circuits are largely absent. Whole-organism models like C. elegans provide a unique advantage: being naturally bacterivorous, worms can be fed live bacterial strains, allowing direct assessment of microbial impacts on protein aggregation, neuronal health and functional endpoints such as motility and behaviour.19 This capability enables studies of gut–-brain interactions that remain inaccessible in even the most advanced in vitro systems.
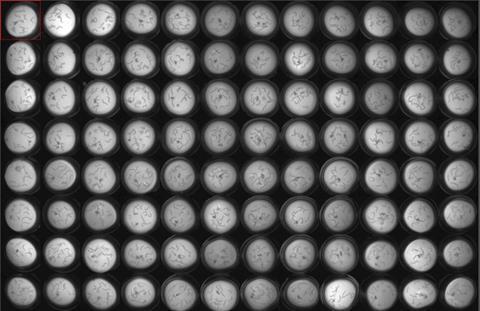

Beyond bridging 3D models and mammalian studies, C. elegans is particularly suited to early target validation and high-throughput drug discovery. Its small size, short lifespan and high reproductive rate make it possible to test hundreds to thousands of compounds efficiently in multiwell formats, linking cellular biomarkers such as protein aggregation with functional readouts like motility, chemotaxis and behavioural assays. Transgenic strains expressing human disease proteins or patient-specific mutations allow modelling of monogenic and polygenic neurodegenerative diseases, while genetic and chemical screens can reveal novel therapeutic targets or repurposing opportunities.20,21
Whole-organism RNA sequencing further enables drug elucidation,22 extending beyond target identification/validation to studying off-target effects and alternative pathways modulated by candidate compounds. This is particularly valuable in neurodegeneration, where interactions between ageing, proteostasis, inflammation and metabolism determine disease progression. By integrating HTS with mechanistic insights at the organism level, C. elegans provides a comprehensive platform for early-stage discovery, functional validation and rational chemical optimisation – all within a scalable, ethically acceptable framework.
Incorporating simple organism models into a tiered non-clinical strategy complements the ongoing development of organoids, microfluidics and AI-driven predictive tools. Rather than aiming to fully replace mammalian models, C. elegans and other lower organisms can reduce reliance on higher animals while capturing critical disease complexity, functional endpoints, systemic interactions and age-dependent pathology that remain inaccessible in vitro. Together, these complementary approaches enable a continuum of discovery – from mechanistic cellular insights to organism-level functional validation – strengthening translational confidence and prioritising candidates with the highest likelihood of clinical success.
Ultimately, a diversified modelling strategy – spanning advanced 3D in vitro systems, perfused co-cultures and ethically tractable whole-organism models – may finally begin to close the translational gap that has stymied decades of neurodegeneration research. While organoids and microfluidics improve physiological fidelity and facilitate studies impossible in 2D, invertebrate models like C. elegans provide a bridge to functional, systemic and ageing-related endpoints. By integrating these platforms with high-throughput and AI-guided discovery, pharma can develop more predictive pipelines, reduce attrition and accelerate therapies that truly improve patient outcomes.
References
1. Cummings J, et al. Alzheimer’s disease drug development pipeline: 2024. Alzheimers Dement (N Y). 2024;10:e12418.
2. Avgerinos KI, et al. Effects of monoclonal antibodies against amyloid-β on clinical and biomarker outcomes and adverse event risks: A systematic review and meta‑analysis of phase III randomized controlled trials in Alzheimer’s disease. Eur J Neurol. 2021;28(6):e26–e39.
3. Zhang Y, et al. Amyloid β‑based therapy for Alzheimer’s disease: challenges and future directions. Signal Transduction and Targeted Therapy. 2023;8(1):177.
4. Duyckaerts C, et al. Models of Alzheimer’s disease: Plaques, tangles, and beyond. Prog Neurobiol. 2015;124:1–18.
5. Mullins C, et al. Toward physiological relevance in organoid models for neurodegenerative diseases. Bioengineering (Basel). 2023;10(3):550.
6. Whitehouse P, et al. Challenges in 3D neural models for neurodegenerative drug discovery. Exp Neurol. 2023;360:114283.
7. Cakir B, et al. Microfluidic models of the blood‑brain barrier. Biotechnol Bioeng. 2017;114(1):184–193.
8. Kim HJ, et al. Human gut‑on‑a‑chip inhabited by microbial flora that experiences intestinal peristalsis‑like motions and flow. Lab Chip. 2012;12:2165–2174.
9. Pamies D, et al. Advanced in vitro models for neurodegeneration research: opportunities and challenges. Nat Rev Neurol. 2021;17(9):558–573.
10. Maoz BM, et al. A linked organ‑on‑chip model of the human neurovascular unit reveals the metabolic coupling of endothelial cells and neurons. Nat Biomed Eng. 2018;2:578–591.
11. UK Government. Replacing animals in science: A strategy to support the development, validation and uptake of non‑animal methods. Department for Science, Innovation and Technology; Nov 2025. Available from: https://www.gov.uk/government/publications/replacing-animals-in-science-strategy/replacing-animals-in-science-a-strategy-to-support-the-development-validation-and-uptake-of-alternative-methods
12. Smith AJ, et al. The UK’s strategy for replacing animal research with predictive biology: implications for translational science. ALTEX. 2026;43(1):12–25.
13. Chen H, et al. Caenorhabditis elegans models for protein aggregation diseases. BMC Chem Biol. 2015;15:14.
14. Roussos P, et al. C. elegans as a model for neurodegenerative diseases. Cells. 2023;12(3):378.
15. Cryan JF, et al. The gut microbiota in neurological disorders. Lancet Neurol. 2020;19(2):179–194.
16. Sampson TR, et al. Gut microbial regulation of motor deficits and neuroinflammation in a model of Parkinson’s disease. Cell. 2016;167(6):1469–1480.e12.
17. Vogt NM, et al. Gut microbiome alterations in Alzheimer’s disease. Sci Rep. 2017;7:13537.
18. Wu S, et al. Leaky gut, endotoxemia and gut microbiota in neurodegenerative diseases. J Neuroinflammation. 2021;18(1):172.
19. Cogliati S, et al. Bacillus Subtilis Delays Neurodegeneration and Behavioral Impairment in the Alzheimer’s Disease Model Caenorhabditis Elegans. J Alzheimers Dis. 2020;73(3):1035-1052.
20. Sohrabi SS, et al. High‑throughput behavioural screen reveals Parkinson’s drug candidates in C. elegans. Commun Biol. 2021;4:597.
21. O’Brien TJ, et al. High‑throughput tracking enables systematic phenotyping and drug repurposing in Caenorhabditis elegans disease models. eLife. 2025;12:e92491.
22. Dwyer DS, et al. Drug elucidation: invertebrate genetics sheds new light on the molecular targets of CNS drugs. Front Pharmacol. 2014;5:177.


















No comments yet