Researchers develop new retinal imaging technique
Posted: 3 January 2017 | Niamh Marriott (Drug Target Review) | No comments yet
Researchers at the University of Rochester Medical Centre have developed a new imaging technique to better assess eye health and potential disease.
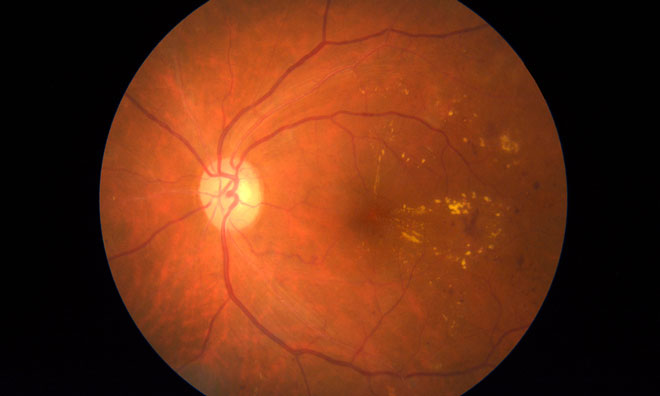

The new technology can make out individual cells in the retina at the back of the eye, which are implicated in vision loss in diseases like glaucoma. They hope their new technique could prevent vision loss via earlier diagnosis and treatment for these diseases.
The group was able to distinguish individual retinal ganglion cells (RGCs), which bear most of the responsibility of relaying visual information to the brain.
Imaging RGCs
There has been a longstanding interest in imaging RGCs because their death causes vision loss in glaucoma, the second leading cause of acquired blindness worldwide. Despite great efforts, no one has successfully captured images of individual RGCs, in part because they are nearly perfectly transparent.
Instead of imaging RGCs directly, glaucoma is currently diagnosed by assessing the thickness of the nerve fibres projecting from the RGCs to the brain. However, by the time retinal nerve fibre thickness has changed detectably, a patient may have lost 100,000 RGCs or more.
“You only have 1.2 million RGCs in the whole eye, so a loss of 100,000 is significant,” said group leader David Williams PhD, University of Rochester, “The sooner we can catch the loss, the better our chances of halting disease and preventing vision loss.”
Ethan Rossi, PhD, University of Pittsburgh School of Medicine, and his colleagues were able to see RGCs by modifying an existing technology – confocal adaptive optics scanning light ophthalmoscopy (AOSLO). They collected multiple images, varying the size and location of the detector they used to gather light scattered out of the retina for each image, and then combined those images.
Multi-offset detection
The technique, called multi-offset detection, was performed at the University of Rochester Medical Centre in animals as well as volunteers with normal vision and patients with age-related macular degeneration.
Not only did this technique allow the group to visualise individual RGCs, but structures within the cells, like nuclei, could also be distinguished in animals. If Rossi can achieve that level of resolution in humans, he hopes to be able to assess glaucoma before the retinal nerve fibre thins – and even before any RGCs die – by detecting size and structure changes in RGC cell bodies.
Cone photoreceptors
While RGCs were the main focus of Rossi’s investigations, they are just one type of cell that can be imaged using this new technique. In age-related macular degeneration, cone photoreceptors that detect colour and are important for central vision are the first to die.
AOSLO has been used to image cones before, but these cells were difficult to see in areas near Drusen, fatty deposits that are the most common early sign of the disease. Using their multi-offset technique in age-related macular degeneration patients, Rossi was able to assess the health of cones near Drusen and in areas where the retina had been damaged.
“This technique offers the opportunity to evaluate many cell classes that have previously remained inaccessible to imaging in the living eye,” said Rossi. “Not only RGCs, but potentially other translucent cell classes and cellular structures.”
Small data
Rossi and his colleagues warn that their study included a small number of volunteers and an even smaller number of age-related macular degeneration patients. More studies will be needed to improve the robustness of the technique and ensure their results are reproducible before it can be widely used in the clinic.
Rossi is now setting up his own laboratory at the University of Pittsburgh and plans to continue working with Williams’ group in studying this technique and its ability to detect changes in retinal cells over the course of retinal diseases.
Related topics
Ophthalmoscopy
Related conditions
Glaucoma, Macular degeneration
Related organisations
Rochester University, University of Pittsburgh School of Medicine
Related people
David Williams, Ethan Rossi



