Enzymes: Ubiquitin-specific proteases as druggable targets
Posted: 21 September 2015 | Anton Simeonov, Mindy Davis (National Institutes of Health)
In much the same way that kinases and phosphatases attach and remove phosphate groups from proteins to modulate their activity, there are a series of enzymes (E1, E2, E3) that add one or more ubiquitins onto a protein, as well as enzymes that remove them (deubiquitinases; DUBs), thereby regulating their activity, location and/or rate of degradation. For various reasons, over time, there has been an increasing interest in targeting the DUBs. The ubiquitin-specific proteases (USPs) comprise the largest family of DUBs, with approximately 56 members in humans, and, since they hold great promise in drug discovery, are the focus of this review.

Ubiquitin is a protein made up of 76 amino acids and is added onto lysines in the target protein through the C-terminal residue of ubiquitin: one can be added (monoubiquitination) or as many as 10 can be added (polyubiquitination). For polyubiquitinated proteins these can be linear or branched chains of ubiquitin, with the complexity of branching reminiscent of the complexity of protein glycosylation. Ubiquitin contains seven lysine residues (Lys6, Lys11, Lys27, Lys29, Lys33, Lys48 and Lys63), and the N-terminal methionine that can be ubiquitinated.
Common linkages include isopeptide bonds through the K48 and K63 on ubiquitin. K48 polyubiquitinated proteins are often targeted to the proteasome for protein degradation and recycling of the ubiquitin. Ubiquitination of a protein can also control its activity/function, such as K63 linkages that regulate the DNA damage response or cell signalling.
Interest in the ubiquitin-proteasome system (UPS) as a target for the treatment of diseases such as cancer, neurodegeneration and autoimmune disease has increased steadily since the approval of the proteasome inhibitors Velcade® (bortezomib) and Kyprolis® (carfilzomib)4. These drugs are used to treat haematological malignancies such as multiple myeloma and mantle cell lymphoma, however, as of yet this drug class has not been approved for solid tumours. Over time, resistance has begun to be observed for this class as well as side effect concerns; this is why there has been an increasing interest in targeting enzymes upstream of the UPS, such as the deubiquitinases (DUBs) and the E3 ligases, which may offer the possibility of greater selectivity and fewer side effects5.
Deubiquitinases
DUBs are situated upstream of the proteasome. The approximately 100 DUB enzymes can be grouped into five main classes, comprising the cysteine proteases ubiquitin C-terminal hydrolases (UCHs), ubiquitin-specific proteases (USPs), ovarian tumour proteases (OTUs), Machado-Joseph domain proteases (MJDs) and the metalloproteases JAB1/MPN/MOV34 (JAMM)6.
As mentioned, the USPs are the largest family of DUBs and are being studied as targets for drug discovery. However, there is still much basic biology to be uncovered for this class of enzymes. Questions surrounding substrate specificity, DUB redundancy and linkage selectivity have yet to be fully addressed for the majority of the class. To date, both linkage-selective DUBs, such as Cezanne, which is specific for Lys11 linkages7, and non-selective DUBs, such as USP2 (which can cleave K48, K63 and linear linkages) have been identified6,8-11. As has been seen for the kinase field12, there is likely room for both selective and nonselective inhibitors as drugs and tool compounds.
The catalytic site of USPs contain a triad with a catalytic cysteine and nearby histidine and asparagine/aspartate to help poise the cysteine for nucleophilic attack. In addition to a USP domain, various USPs have additional domains, such as ubiquitin-like domains and zinc-finger domains6. Additionally, several of the USPs function as complexes, such as USP1/UAF1, USP12/UAF1/WDR20 and USP46/UAF1/WDR2013,14. Several USPs have crystal structures reported in the PDB, including USP2 (PDB ID 2HD5), USP5 (PDB ID 3IHP), USP7 (PDB ID 4M5W), USP14 (PDB ID 2AYN), CYLD (PDB ID 2VHF) and USP21 (2Y5B). In an analogous way to kinases, USPs also seem to have active and inactive conformations with active conformations observed upon ubiquitin binding, although also like kinases not every USP has been observed in both conformations6,15.
Assay technologies to interrogate DUBs
In order to identify DUB inhibitors, DUB substrates and DUB inhibitor selectivity, a variety of assay reagents have been identified and utilised in high-throughput screening (HTS) campaigns as well as lower throughput gel and western blot experiments (Figure 1)16-18. The higher throughput methods generally involve an increase in luminescence or fluorescence upon cleavage that can be monitored on a plate reader16. Commonly-used reagents are ubiquitin linked to a fluorophore through a linear linkage, such as Ub-AMC (Ub-7-amino-4-methylcoumarin) and Ub-Rhodamine110 (Figure 1A), which have been used for screening various USPs including USP1 (PubChem Assay Identifier (AID) 504865), USP2 (PubChem AID 493170) and USP14 (PubChem AID 449747).
More recently, reagents have been created that contain an isopeptide linkage between a di-ubiquitin (Di-Ub) to more closely mimic the most common in vivo Ub linkage. One example of this type of assay involves using an internally quenched fluorescent reagent in which one Ub has a fluorophore and the other has a quencher that quenches the fluorophore when the two are in close proximity but not once the Di-Ub is cleaved (Figure 1B) 8. Another method that has been utilised represents a coupled enzyme system. In one format, called Ub-Chop2, the ubiquitin is linked to an enzyme that is only active when released and thereby can produce a fluorescence enzyme product (Figure 1C); Ub-Chop2 has been used to identify inhibitors of USP2 in a large-scale screen (PubChem AID 463254).
Additionally, having these reagents with different fluorescence excitation and emission wavelengths can help mitigate compound interference, which can be particularly problematic in the blue spectral region19, such as for Ub-AMC, by allowing for orthogonal assay development. Lastly, when aminoluciferin is tethered to the DUB substrate Z-RLRGG, it is not luminescent but upon cleavage by a DUB, its luminescence can be measured by a luminometer (Figure 1D), a technique that was used for USP820. All of these reagents can be read out in kinetic mode which can help minimize assay interference from compound fluorescence. Additional methods for identifying DUB substrates, DUB inhibitors and DUB linkage preferences include mass spectrometry-based methods and protein microarray methods17,18,21.
Lower throughput methods that are important for confirmation of hits rely primarily on a change in mass upon Di-Ub cleavage or DUB binding. Ubiquitin aldehyde or ubiquitin vinyl sulfone are reagents that form covalent irreversible linkages to DUBs upon binding, leading to observable mass changes to the DUBs (Figure 1E). These have been visualised in western blots with antibodies to the individual DUB, such as USP1, or to ubiquitin, or to a tag, such as HA, placed on the ubiquitin, HA-ubiquitin vinyl sulfone22. Additionally, Di-Ub can be cleaved by a DUB and this new product can be visualised on a gel as a band with half the molecular weight of the substrate (Figure 1F), a technique that has been used for USP123. Tetra-ubiquitin molecules are becoming increasingly available that can also be used for this purpose. Having a diversity of assay technologies is important for identifying and validating inhibitors and studying the USP family.
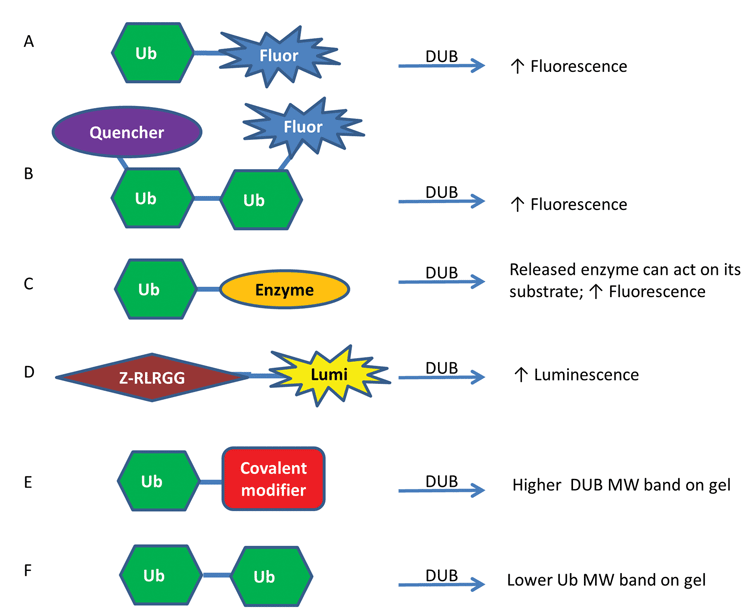

Figure 1. Representative assay formats used to study DUBs
USPs as drug targets
While the USP family as a whole is still widely unexplored, selected USPs have been the focus of intense research efforts both to uncover their physiological roles and to identify inhibitors24,25, with some of the highlights to date described below.
USP1
USP1 deubiquitinates PCNA and FANCD2, which are proteins important for DNA repair pathways and the Fanconi Anemia pathway, respectively3. USP1, which is active as a complex with UAF113, can deubiquitinate PCNA, which upon DNA damage is ubiquitinated and thereby recruits translesion DNA polymerase eta. ML323 was developed as an inhibitor of USP1/UAF1 following a high-throughput screen using Ub-Rhodamine110 assay26. The additional compounds pimozide, GW7647 and C527, were found to be sub µM inhibitors22,23.
USP2
USP2, found in vivo as the alternately spliced USP2a and USP2b, has a large number of identified substrates important in cancer including cyclin D1, MDM2, and fatty acid synthase (FAS)27-29. Cyclin D1, which can be overexpressed in cancer, was degraded in response to USP2-targeting siRNA leading to inhibition of cell cycle progression in cancer cells but not normal cells28. USP2 can also deubiquitinate MDM2, thereby destabilising p5329. USP2a was shown to help protect prostate cancer cells from apoptosis by deubiquitinating the antiapoptotic proteins MDM2 and FAS, which can be overexpressed in cancer27,30. Indeed 44% of prostate tumours tested had overexpression of USP2a. 2-cyano-pyrimidines and -triazines have been identified as inhibitors of USP2 and UCH-L331.
USP5
USP5 has been implicated in suppression of both p53 and FAS levels in melanoma cells. An inhibitor, EOAI3402143, which had been optimised from WP113032, was shown to recapitulate USP5 knockdown results and block melanoma growth in a mouse model33. Inhibition was able to overcome resistance to BRAF-targeting kinase inhibitors in melanoma33. This compound also inhibits USP9x, a drug target discussed below34-36.
USP7/HAUSP
USP7/HAUSP (herpes virus-associated ubiquitin-specific protease) deubiquitinates MDM2 (an E3 ligase; also called HDM2), thereby destabilising p53 and can be targeted by small molecule inhibitors37. Additional roles for USP7/HAUSP include deubiquitination of the tumour suppressors PTEN (phosphatase and tensin homologue deleted in chromosome 10) and FOXO4 (Forkhead box O), which favours their localisation to the cytoplasm versus the nucleus limiting their transcriptional activity38,39. It should be noted that PTEN is also deubiquitinated by USP1340. USP7/HAUSP is overexpressed in cancer, such as prostate cancer. Inhibitors of USP7 include HBX 41,108 and P22077, which were identified from HTS campaigns37,41,42.
USP8
USP8 (UBPY) knockdown in gefitinib-resistant non-small cell lung cancer (NSCLC) but not gefitinib-sensitive NSCLC leads to cell death, potentially providing an avenue to pursue when resistance to EGFR receptor kinase inhibitors develops20. This knockdown effect could be recapitulated with a USP8 inhibitor (9-ethyloxyimino-9H-indeno[1,2-b]pyrazine-2,3-dicarbonitrile)43 which showed efficacy in a mouse xenograft model. USP8 had previously been shown to be involved in EGFR degradation44. Additionally, USP8 mutations cause Cushing’s Disease, a disease caused by pituitary corticotroph adenomas hypersecting adenocorticotropin (ACTH)45. The mutations cause a decrease in binding to 14-3-3 protein, increased cleavage of USP8 and an increase in DUB activity, which leads to increased recycling of EGFR, thus raising the levels of ACTH 45.
USP9x
USP9x plays an important role in stabilising beta-catenin35, MCL136 and SMAD434, a component of the TGFβ signaling pathway. Each of these proteins is in its own right a target for cancer therapy. USP9x is one of the DUBs inhibited by WP113032.
USP12
USP12 is active as a complex (USP12/UAF1/WDR20) and has several substrates identified to date including the androgen receptor, which is important in prostate and some breast cancers14,46. To date, only one compound has been reported to inhibit USP12/UAF1/WDR20 and that is GW7647, which also inhibits USP123,47.
USP14
USP14 is associated with the proteasome and is one of the DUBs responsible for ensuring that ubiquitin is recycled rather than degraded by the proteasome, contributing to ubiquitin homeostasis. An inhibitor of USP14 was identified and shown to enhance the activity of the proteasome48. Inhibition of USP14 lead to accelerated proteasomal degradation of proteins involved in neurodegenerative diseases.
USP28
USP28 has a role in stabilising the oncoprotein c-myc as discovered in a shRNA screen49, but is also implicated in other functions that may lead to side effects5. For example, USP28 is important for the DNA damage response50. USP28 is highly expressed in colon and breast carcinomas and has been implicated in conferring stem-cell-like traits to breast cancer cells51.
Perspective
DUBs have been a subject of increased interest as a potential novel drug target class of late, in turn spurring efforts to develop relevant activity assays. Most reagents used for DUB assays contain one Ub linked to a fluorophore or luminescent compound by a linear linkage or two Ubs linked by an isopeptide linkage, but polyubiquitinated proteins can have as many as 10 Ubs linked together in linear or branched conformations1,9. Having additional assay reagents, such as tetraubiquitin, with >2 ubiquitins as well as branched Ubs, may be important for furthering the understanding of Dub linkage selectivity. It will be interesting to see the impact on HTS campaigns of utilising the different available assay reagents to understand whether specific inhibitors exist targeting the different linkages and poly-ubiquitin reagents.
Screening a large panel size was important to obtain an accurate assessment of inhibitor selectivity for kinases12, and it will be interesting to see whether the same will hold for DUB families, such as the USPs, or whether a small panel will accurately represent overall selectivity. There is a need for the development and execution of HTS assays and the discovery of molecule probes for all of the ~56 USPs in order to understand their biology. Once a critical mass of inhibitors are available, a broad selectivity profile will be important in order to understand whether highly selective modulators across the whole DUB class can be obtained as is the case for some kinase inhibitors, such as lapatinib for EGFR, which is selective across the whole kinome12. A given DUB, such as USP2, can have multiple substrates that it can deubiquitinate, so in addition to polypharmacology from inhibitors that hit more than one DUB, even a DUB-selective inhibitor may impact multiple cellular proteins and processes due to multiple substrates acted on in vivo.
Additionally, redundancy within the USP family is already being observed with multiple DUBs impacting MDM2, p53, FAS and PTEN27,29,33,37,38. Thus, the DUB field seems poised for rapid development: in much the same way as the approval of Gleevec sparked fervor into the kinase field52, approval of a DUB inhibitor would be expected to do the same for the deubiquitinase field.
References
- Komander D, Rape M. The ubiquitin code. Annu Rev Biochem. 2012;81:203-29
- Moremen KW, Tiemeyer M, Nairn AV. Vertebrate protein glycosylation: diversity, synthesis and function. Nat Rev Mol Cell Biol. 2012;13(7):448-62
- Jacq X, Kemp M, Martin NM, Jackson SP. Deubiquitylating enzymes and DNA damage response pathways. Cell Biochem Biophys. 2013 Sep;67(1):25-43
- Dou QP, Zonder JA. Overview of proteasome inhibitor-based anti-cancer therapies: perspective on bortezomib and second generation proteasome inhibitors versus future generation inhibitors of ubiquitin-proteasome system. Curr Cancer Drug Targets. 2014;14(6):517-36
- Shi D, Grossman SR. Ubiquitin becomes ubiquitous in cancer: emerging roles of ubiquitin ligases and deubiquitinases in tumorigenesis and as therapeutic targets. Cancer Biol Ther. 2010 Oct 15;10(8):737-47
- Komander D, Clague MJ, Urbe S. Breaking the chains: structure and function of the deubiquitinases. Nat Rev Mol Cell Biol. 2009 Aug;10(8):550-63
- Bremm A, Freund SMV, Komander D. Lys11-linked ubiquitin chains adopt compact conformations and are preferentially hydrolyzed by the deubiquitinase Cezanne. Nat Struct Mol Biol. 2010;17(8):939-47
- Chelur D, Eddins M, Leach C, Orcutt SJ, Peroutka RJ, Strickler J, et al., inventors; Progenra, Inc., Lifesensors, Inc., assignee. Di- and poly-ubiquitin deubiquitinase substrates and uses thereof2013
- Faesen AC, Luna-Vargas MP, Geurink PP, Clerici M, Merkx R, van Dijk WJ, et al. The differential modulation of USP activity by internal regulatory domains, interactors and eight ubiquitin chain types. Chem Biol. 2011 Dec 23;18(12):1550-61
- Komander D, Reyes-Turcu F, Licchesi JDF, Odenwaelder P, Wilkinson KD, Barford D. Molecular discrimination of structurally equivalent Lys 63-linked and linear polyubiquitin chains. EMBO Reports. 2009;10(5):466-73
- Nijman SM, Luna-Vargas MP, Velds A, Brummelkamp TR, Dirac AM, Sixma TK, et al. A genomic and functional inventory of deubiquitinating enzymes. Cell. 2005 Dec 2;123(5):773-86
- Davis MI, Hunt JP, Herrgard S, Ciceri P, Wodicka LM, Pallares G, et al. Comprehensive analysis of kinase inhibitor selectivity. Nat Biotechnol. 2011 Nov;29(11):1046-51
- Cohn MA, Kowal P, Yang K, Haas W, Huang TT, Gygi SP, et al. A UAF1-containing multisubunit protein complex regulates the Fanconi anemia pathway. Molecular cell. 2007 Dec 14;28(5):786-97
- Kee Y, Yang K, Cohn MA, Haas W, Gygi SP, D’Andrea AD. WDR20 Regulates Activity of the USP12·UAF1 Deubiquitinating Enzyme Complex. Journal of Biological Chemistry. 2010 April 9, 2010;285(15):11252-7
- Nolen B, Taylor S, Ghosh G. Regulation of Protein Kinases: Controlling Activity through Activation Segment Conformation. Molecular Cell. 2004;15(5):661-75
- Kramer HB, Nicholson B, Kessler BM, Altun M. Detection of ubiquitin–proteasome enzymatic activities in cells: Application of activity-based probes to inhibitor development. Biochimica et Biophysica Acta (BBA) – Molecular Cell Research. 2012;1823(11):2029-37
- Loch CM, Strickler JE. A microarray of ubiquitylated proteins for profiling deubiquitylase activity reveals the critical roles of both chain and substrate. Biochimica et Biophysica Acta (BBA) – Molecular Cell Research. 2012;1823(11):2069-78
- Ritorto MS, Ewan R, Perez-Oliva AB, Knebel A, Buhrlage SJ, Wightman M, et al. Screening of DUB activity and specificity by MALDI-TOF mass spectrometry. Nat Commun. 2014;5:4763
- Simeonov A, Jadhav A, Thomas CJ, Wang Y, Huang R, Southall NT, et al. Fluorescence spectroscopic profiling of compound libraries. J Med Chem. 2008 Apr 24;51(8):2363-71
- Byun S, Lee SY, Lee J, Jeong CH, Farrand L, Lim S, et al. USP8 is a novel target for overcoming gefitinib resistance in lung cancer. Clin Cancer Res. 2013 Jul 15;19(14):3894-904
- McGouran JF, Gaertner SR, Altun M, Kramer HB, Kessler BM. Deubiquitinating enzyme specificity for ubiquitin chain topology profiled by di-ubiquitin activity probes. Chem Biol. 2013 Dec 19;20(12):1447-55
- Mistry H, Hsieh G, Buhrlage SJ, Huang M, Park E, Cuny GD, et al. Small-molecule inhibitors of USP1 target ID1 degradation in leukemic cells. Mol Cancer Ther. 2013 Dec;12(12):2651-62
- Chen J, Dexheimer TS, Ai Y, Liang Q, Villamil MA, Inglese J, et al. Selective and cell-active inhibitors of the USP1/ UAF1 deubiquitinase complex reverse cisplatin resistance in non-small cell lung cancer cells. Chem Biol. 2011 Nov 23;18(11):1390-400
- Edelmann MJ, Nicholson B, Kessler BM. Pharmacological targets in the ubiquitin system offer new ways of treating cancer, neurodegenerative disorders and infectious diseases. Expert Rev Mol Med. 2011;13:e35
- Farshi P, Deshmukh RR, Nwankwo JO, Arkwright RT, Cvek B, Liu J, et al. Deubiquitinases (DUBs) and DUB inhibitors: a patent review. Expert Opin Ther Pat. 2015 Jun 16:1-18
- Liang Q, Dexheimer TS, Zhang P, Rosenthal AS, Villamil MA, You C, et al. A selective USP1–UAF1 inhibitor links deubiquitination to DNA damage responses. Nat Chem Biol. 2014;10(4):298-304
- Graner E, Tang D, Rossi S, Baron A, Migita T, Weinstein LJ, et al. The isopeptidase USP2a regulates the stability of fatty acid synthase in prostate cancer. Cancer Cell. 2004;5(3):253-61
- Shan J, Zhao W, Gu W. Suppression of Cancer Cell Growth by Promoting Cyclin D1 Degradation. Molecular Cell. 2009;36(3):469-76
- Stevenson LF, Sparks A, Allende‐Vega N, Xirodimas DP, Lane DP, Saville MK. The deubiquitinating enzyme USP2a regulates the p53 pathway by targeting Mdm2. The EMBO Journal. 2007;26(4):976-86#
- Priolo C, Tang D, Brahamandan M, Benassi B, Sicinska E, Ogino S, et al. The isopeptidase USP2a protects human prostate cancer from apoptosis. Cancer Res. 2006 Sep 1;66(17):8625-32
- Flohr S, Furet P, Imbach P, Hommel U, Litscher HU, Gil PS, et al., inventors; 2-cyano-pyrimidines and -triazines as cysteine protease inhibitors; 2007
- Kapuria V, Peterson LF, Fang D, Bornmann WG, Talpaz M, Donato NJ. Deubiquitinase inhibition by small-molecule WP1130 triggers aggresome formation and tumor cell apoptosis. Cancer Res. 2010 Nov 15;70(22):9265-76
- Potu H, Peterson LF, Pal A, Verhaegen M, Cao J, Talpaz M, et al. Usp5 links suppression of p53 and FAS levels in melanoma to the BRAF pathway. Oncotarget. 2014 Jul 30;5(14):5559-69
- Dupont S, Mamidi A, Cordenonsi M, Montagner M, Zacchigna L, Adorno M, et al. FAM/USP9x, a Deubiquitinating Enzyme Essential for TGFβ Signaling, Controls Smad4 Monoubiquitination. Cell. 2009;136(1):123-35
- Murray RZ, Jolly LA, Wood SA. The FAM Deubiquitylating Enzyme Localizes to Multiple Points of Protein Trafficking in Epithelia, where It Associates with E-cadherin and β-catenin. Molecular Biology of the Cell. 2004 April 1, 2004;15(4):1591-9
- Schwickart M, Huang X, Lill JR, Liu J, Ferrando R, French DM, et al. Deubiquitinase USP9X stabilizes MCL1 and promotes tumour cell survival. Nature. 2010;463(7277):103-7
- Colland F, Formstecher E, Jacq X, Reverdy C, Planquette C, Conrath S, et al. Small-molecule inhibitor of USP7/HAUSP ubiquitin protease stabilizes and activates p53 in cells. Mol Cancer Ther. 2009 Aug;8(8):2286-95
- Song MS, Salmena L, Carracedo A, Egia A, Lo-Coco F, Teruya-Feldstein J, et al. The deubiquitinylation and localization of PTEN are regulated by a HAUSP-PML network. Nature. 2008;455(7214):813-7
- van der Horst A, de Vries-Smits AMM, Brenkman AB, van Triest MH, van den Broek N, Colland F, et al. FOXO4 transcriptional activity is regulated by monoubiquitination and USP7/HAUSP. Nat Cell Biol. 2006;8(10):1064-73
- Zhang J, Zhang P, Wei Y, Piao HL, Wang W, Maddika S, et al. Deubiquitylation and stabilization of PTEN by USP13. Nature cell biology. 2013 Dec;15(12):1486-94
- Reverdy C, Conrath S, Lopez R, Planquette C, Atmanene C, Collura V, et al. Discovery of specific inhibitors of human USP7/HAUSP deubiquitinating enzyme. Chem Biol. 2012 Apr 20;19(4):467-77
- Weinstock J, Wu J, Cao P, Kingsbury WD, McDermott JL, Kodrasov MP, et al. Selective Dual Inhibitors of the Cancer-Related Deubiquitylating Proteases USP7 and USP47. ACS Med Chem Lett. 2012 Oct 11;3(10):789-92
- Colombo M, Vallese S, Peretto I, Jacq X, Rain JC, Colland F, et al. Synthesis and biological evaluation of 9-oxo-9H-indeno[1,2-b]pyrazine-2,3-dicarbonitrile analogues as potential inhibitors of deubiquitinating enzymes. ChemMedChem. 2010 Apr 6;5(4):552-8
- Alwan HAJ, van Leeuwen JEM. UBPY-mediated Epidermal Growth Factor Receptor (EGFR) De-ubiquitination Promotes EGFR Degradation. Journal of Biological Chemistry. 2007;282(3):1658-69
- Reincke M, Sbiera S, Hayakawa A, Theodoropoulou M, Osswald A, Beuschlein F, et al. Mutations in the deubiquitinase gene USP8 cause Cushing’s disease. Nat Genet. 2015;47(1):31-8
- Burska UL, Harle VJ, Coffey K, Darby S, Ramsey H, O’Neill D, et al. Deubiquitinating Enzyme Usp12 Is a Novel Co-activator of the Androgen Receptor. Journal of Biological Chemistry. 2013 November 8, 2013;288(45):32641-50
- McClurg UL, Summerscales EE, Harle VJ, Gaughan L, Robson CN. Deubiquitinating enzyme Usp12 regulates the interaction between the androgen receptor and the Akt pathway. Oncotarget. 2014 Aug 30;5(16):7081-92
- Lee BH, Lee MJ, Park S, Oh DC, Elsasser S, Chen PC, et al. Enhancement of proteasome activity by a small-molecule inhibitor of USP14. Nature. 2010 Sep 9;467(7312):179-84
- Popov N, Wanzel M, Madiredjo M, Zhang D, Beijersbergen R, Bernards R, et al. The ubiquitin-specific protease USP28 is required for MYC stability. Nat Cell Biol. 2007;9(7):765-74
- Zhang D, Zaugg K, Mak TW, Elledge SJ. A Role for the Deubiquitinating Enzyme USP28 in Control of the DNA-Damage Response. Cell. 2006;126(3):529-42
- Wu Y, Wang Y, Yang XH, Kang T, Zhao Y, Wang C, et al. The deubiquitinase USP28 stabilizes LSD1 and confers stem-cell-like traits to breast cancer cells. Cell Rep. 2013 Oct 17;5(1):224-36
- Cohen MH, Williams G, Johnson JR, Duan J, Gobburu J, Rahman A, et al. Approval Summary for Imatinib Mesylate Capsules in the Treatment of Chronic Myelogenous Leukemia. Clinical Cancer Research. 2002 May 1, 2002;8(5):935-42
Biographies
Mindy I. Davis is a Senior Research Scientist in the Division of Preclinical Innovation at National Center for Advancing Translational Sciences (NCATS), part of the NIH. Prior to NCATS, she was a Senior Research Scientist at Ambit Biosciences, focused on assay development and high-throughput screening. Her chemistry PhD was obtained from Stanford and her biology postdoc was at Caltech. She has extensive experience in developing biochemical and cell-based HTS assays across multiple target classes, including deubiquitinases and kinases.
Dr. Simeonov is the Acting Scientific Director at the National Center for Advancing Translational Sciences within the National Institutes of Health. He holds a PhD in bioorganic chemistry from the University of Southern California and a BA in chemistry from Concordia College; he trained as a postdoctoral fellow at The Scripps Research Institute under Professors Richard Lerner and Kim Janda. His research interests include novel detection chemistries and techniques, assays for clinical diagnostics, assay miniaturisation and novel approaches to screening and small molecule development. Dr. Simeonov, the author and inventor on over 130 peer-reviewed scientific publications and patents, has a truly diverse background, ranging from bioorganic chemistry and molecular biology to clinical diagnostic research and development. Prior to joining NIH in November 2004, he was a senior scientist at Caliper Life Sciences, a leading developer of microfluidic technologies, where he was responsible for both basic research on novel assay methodologies and development of microfluidic products for research and clinical diagnostics. Dr. Simeonov is a Literature Editor of Assay and Drug Development Technologies and a member of the Editorial Board of Drug Target Review and Expert Opinion on Drug Discovery.
Related topics
Drug Discovery, Enzymes
Related organisations
National Institutes of Health (NIH)
Related people
Anton Simeonov, Mindy Davis