Enabling new targets by integrated lead discovery
Posted: 3 December 2015 | Sylvain Cottens (Novartis), Ulrich Schopfer (Novartis) | No comments yet
Progress in the understanding of disease mechanisms provides new opportunities to discover molecules that modulate disease. To capitalise on these opportunities, successful lead discovery strategies need to build on insights into how a cellular phenotype or a target contributes to disease biology. Recent advances in induced pluripotent stem cells (iPSCs), gene editing and imaging technologies enable unprecedented access to relevant cellular models of disease, while methods such as fragment-based and DNA-encoded library screening have expanded our ability to find ligands, even for difficult and unusual targets. Realising that only a tight integration of approaches can overcome the challenges of these novel targets, we introduce the concept of ‘integrated lead discovery’ and highlight the scientific, technological and organisational implications of the concept.


The advances that have been made in biomedical understanding of disease need to be translated into treatments to improve the lives of patients. Lead discovery is a key step in this translation, aiming at finding molecules that modulate a target or phenotype to affect their role in disease. However, despite successes, there remains a need to further improve lead discovery, and in particular to address unprecedented and difficult target classes. High-throughput screening (HTS) was widely adopted in the 1990s to tackle this problem. Especially in combination with combinatorial chemistry, HTS was considered a way to turn drug discovery from a game of chances into a rational, predictable industrial process.
Later, HTS was rather seen as a contributor to the escalating research and development (R&D) costs in the pharmaceutical industry without significantly adding to pipelines1,2. It is only recently that a more balanced assessment of the role and contribution of HTS to drug discovery has become possible and today’s R&D pipelines contain many drug candidates that originate from HTS.3 A closer look, however, shows that a drug was rarely discovered by a single technology, and frequently it is difficult to retrospectively trace the relative contributions of screening, design, structural information, in silico modelling or other technologies in the complex and highly iterative path of drug discovery.
Meanwhile, the recent renaissance of phenotypic drug discovery4 has been accompanied by advances in cell biology and imaging technologies that allow the development of more disease-relevant cellular models. Working with these more complex models means that lead discovery becomes even less of a standardised process. This led us to re-think our approach to lead discovery. We saw the need to place the disease-relevant pathway or target at the centre of our thinking. Based on the biological understanding of disease pathology, we then brought the breadth of our capabilities to bear on the scientific hypothesis, with an emphasis on the integration of technologies. We termed this concept “integrated lead discovery (ILD)” (Figure 1).
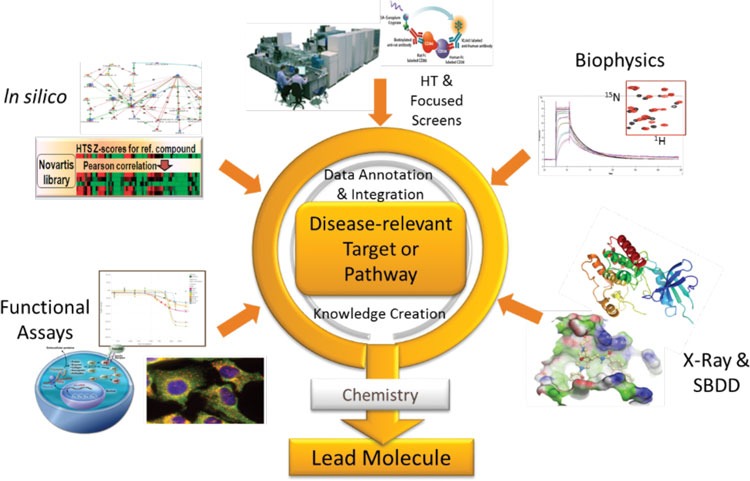

Figure 1: Integrated lead discovery: a project perspective. Synergy between lead discovery approaches
A new concept for lead discovery
ILD has its foundation in the biological understanding of the role of a target or pathway in disease. A thorough understanding of how a cellular phenotype or a signalling pathway contributes to disease can be used to design lead discovery strategies that capture essential elements of biology. Primary cells or iPSC-derived cellular systems play an increasing role here, since they are more reflective of the disease tissue than transformed cell lines. Live or fixed cell imaging with advanced image analysis are powerful methods to interrogate cellular systems. In silico methods help to select focused compound sets and to examine results in the context of pathway knowledge. Similarly, a detailed understanding of the role of a protein complex, a protein, or a specific mechanism of action of a protein domain can be incorporated into strategies that capitalise on a unique insight into protein function, and which are also open to non-canonical findings. Strong protein biochemistry capabilities, in silico modelling and X-ray crystallography combined with biochemical and biophysical assays that detect binding or enzyme activity can build on the strength of each technology.
An essential element of the ILD concept is to also pursue low affinity/low activity hits. To differentiate theses hits from assay noise and false positives, biophysical methods that show true compound binding to the target protein play a critical role. Also, fragment-based discovery efforts that use small, low-affinity molecules to interrogate binding sites on proteins are a core element of ILD, because fragments have a higher likelihood of binding to a protein pocket.
Importantly, the concept implies that we aim to develop the most biologically-relevant assay even when it does not offer the throughput to test millions or even hundreds of thousands of compounds. In most projects, HTS has been replaced by iterations of low- or medium-throughput assays, testing between 1000 to 100,000 compounds at each run. Hits from these runs provide tool compounds that can be used to develop follow-up assays or even to challenge the underlying project hypothesis. The ILD approach requires a tight interaction and close communication across disciplines. As no two projects follow the same process, it is vital that project teams learn from each other’s experiences and accumulate a body of knowledge across projects. These requirements represent significant changes to earlier approaches. To realise the full potential of ILD, a shift in culture and a new organisation were required.
A new culture
As in most other companies, lead discovery at Novartis started in a central department that had established the technologies and processes required to test large numbers of compounds in high-throughput assays. Also for structural biology, a process flow had been developed and partially automated that optimised efficiency at each step to maximise the number of structures that could be solved. This had created a culture of technology experts who took pride in the processes they had developed. This mind-set had to be shifted in favour of emphasising biological relevance. Numerical performance indicators were replaced by assessing how contributions had enabled lead discovery. While performance and innovation were still core values of the organisation, they were complemented by a strong emphasis on collaboration.
Integrating knowledge across disciplines and sharing credit with colleagues, while being easy to work with for partners in disease areas and medicinal chemistry, required training in self-awareness and communication. It was also designed to question the leadership model5 of the organisation. Leaders took the role of mentors, who help with problem-solving and connecting people. They establish the direction but refrain from prescribing a solution. In this way they enable individuals, teams and the entire organisation to learn. As we found out, the standard hierarchical organisational model, with technology-centric groups at its core, was not flexible enough to enable project-centric, fluid teams to form and disassemble over time.
A new organisation
A new matrix organisation was built along two axes: lead discovery projects and technologies (Figure 2). Projects were grouped according to disease area to facilitate interactions with disease areas and project portfolio management. Technologies were grouped to build strong expertise cores. Mid-level group leaders were replaced by a strengthened project leader role. These leaders have the responsibility of developing and executing the project lead discovery strategy, together with the core project team from the disease area and from chemistry. They could come from any discipline but were expected to have sufficient understanding of the other disciplines to be able to guide strategy development.
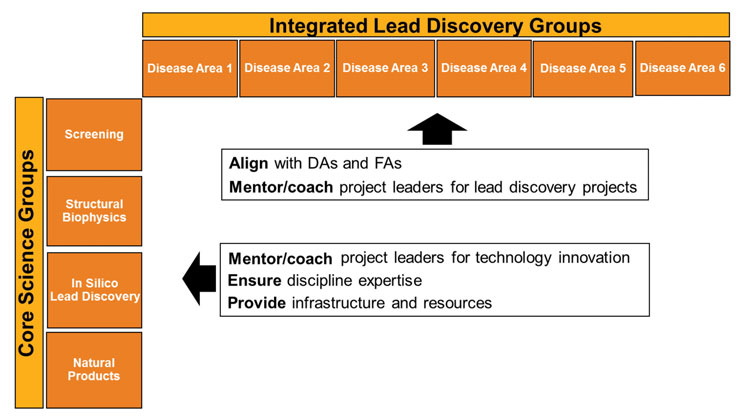

Figure 2: A new lead discovery organization. A matrix with focus on projects and on scientific disciplines
ILD groups united projects into a portfolio that partnered with the same disease area. The leader of this ILD group became a member of the disease area management board under the agreement that this board would be the single governance board for that project. In that role, he or she mentors project leaders and ensures alignment with the disease area with regard to scientific strategy and project prioritisation. Along the second axes, Core Science Groups were formed to ensure the continued evolution of capabilities and skills. The leaders of these groups provide the infrastructure and resources for the projects, as well as guiding innovation projects that advance the technology in their area. They ensure flexibility of resource assignment across the ILD groups as the size and nature of the project portfolio of any disease area changes over time.
The organisation was designed to maximise synergies across technologies and approaches, as well as across project portfolios. Project teams retain a high degree of scientific autonomy to empower them to from strong collaborations with partners in disease areas and chemistry and quickly react to scientific discoveries.
Integrated lead discovery in action
The experience with this model has been overwhelmingly positive. While in the past the lead discovery group was often viewed as slow, bureaucratic and process-driven, the perception has changed and the group has now become a strong, flexible and science-driven partner for disease areas and other platforms. Project teams address key questions in a project by applying relevant expertise and technologies rather than executing a project plan. Rapid iterations between small pilot screens, early hit confirmation, input by early structure and biophysical information have become standard practice. These activities can generate tool compounds to validate flowcharts all the way to providing in vivo concept validation and thus bridge between phenotypic- and target-based approaches.
For targets that were previously deemed ‘undruggable’, activity-agnostic binding assays are used to find ligands, identify binding sites and elucidate mechanisms of action. Fragment-based drug discovery using biophysical methods such as nuclear magnetic resonance, differential scanning fluorimetry or others has proven a powerful approach. Also, new selection-based screening technologies, such as DNA-encoded libraries, are used to gain an understanding of pockets that can be used to modulate target activity. In combination with in silico methods, ligands can rapidly be improved to become higher in affinity and enable a proof of principle in cells (Figure 3).
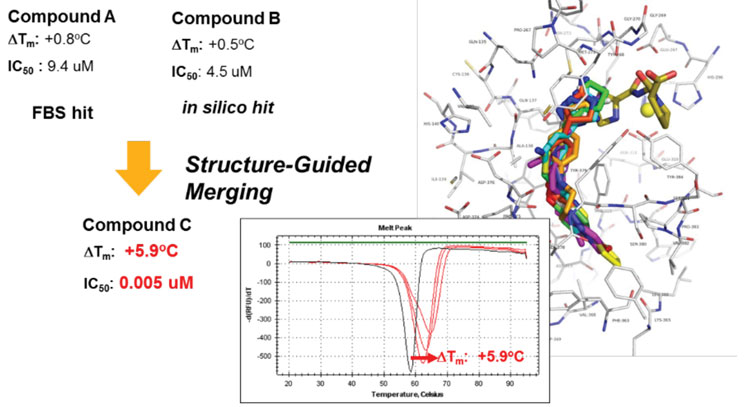

Figure 3: Ligands identified by fragment-based screening and in silico docking were merged and guided by co-crystal structures and biochemical assays to yield potent inhibitors
Initially, a concern was that the project focus in the new organisation would lead to reduced efforts in technology development and that it would become more difficult to dedicate resources to technology innovation. This has not been the case. On the contrary, activities in technological innovation have become more deliberate and more strategic, with a focus on keeping the technology base of the department at the forefront of the discipline.
However, the adaptation to working in a matrix organisation is not without difficulties. Since there is a high degree of co-dependency across the matrix, the organisation needs a high level of trust and communication to function well. Decision-making can be more complex than in a hierarchical organisation. Initially, scientists were not sure who to approach to get decisions, because depending on the topic, they interacted with different leadership team members. The closer collaboration with a disease-area partner in the ILD groups therefore has two sides. On balance, however, the ILD approach has transformed the department to become more flexible and more innovative. Delayering and focused leadership roles strengthened both the project and the technology focus of the organisation and empowered associates to take ownership of the scientific challenge of lead discovery.
Conclusion
Rethinking our approach to lead discovery and the understanding of disease mechanisms has been placed at the centre of lead discovery strategies. Key scientific questions are addressed in an iterative, hypothesis-driven manner. We believe that this flexible, science-driven approach will give rise to the next generation of drug targets for which the reliance on standard flowcharts and processes will no longer be suitable.
References
- Peter Landers, Human Element: drug industry’s big push into technology falls short — testing machines were built to streamline research — but may be stifling it — officials see payoff after 2010. The Wall Street Journal (24 Feb 2004).
- Roger Lahana, Who wants to be irrational? Drug Discov. Today. 2003; 8, 655–656
- Ricardo Macarron, Martyn N. Banks, Dejan Bojanic, David J. Burns, Dragan A. Cirovic, Tina Garyantes, Darren V. S. Green, Robert P. Hertzberg, William P. Janzen, Jeff W. Paslay, Ulrich Schopfer, G. Sitta Sittampalam, Impact of high-throughput screening in biomedical research. Nature Reviews Drug Discovery. 2011; 10, 188-195
- David C. Swinney, Jason Anthony, How were new medicines discovered? Nature Reviews Drug Discovery. 2011; 10, 507-519
- Andreas Schneider, Zeynep Erden, Hans Widmer, Guido Koch, Christine Billy, Georg von Krogh. Rethinking leadership in drug discovery projects. Drug Discovery Today. 2012; 23–24, 1258-1262
Biographies
Ulrich Schopfer is Executive Director and Integrated Lead Discovery Head at the Center of Proteomic Chemistry of the Novartis Institutes for BioMedical Research (NIBR). He is overseeing lead discovery projects in the oncology and muscoskeletal disease area as well as lead discovery projects partnered with the disease molecular pathways group. Previously he led a biochemical screening group and was responsible for global compound management at NIBR. Ulrich Schopfer holds a PhD in chemistry and started his career at Novartis as Medicinal Chemist and Project Team Leader of different discovery programs in Neuroscience.
Sylvain Cottens is the Global Head of the Center for Proteomic Chemistry at the Novartis Institutes for Biomedical Research (NIBR). Previously he led the Novartis Expertise Platform Proteases. Sylvain Cottens holds a PhD in chemistry and started his career at Novartis as a medicinal chemist, first in the Neurosciences area and then in the Immunology department, before becoming a medicinal Chemistry Unit Head. From 2000 to 2002, Sylvain Cottens was the acting head of the Novartis Transplantation Research department.
Related topics
Drug Discovery, High-Throughput Screening (HTS), Screening
Related organisations
Novartis
Related people
Sylvain Cottens, Ulrich Schopfer