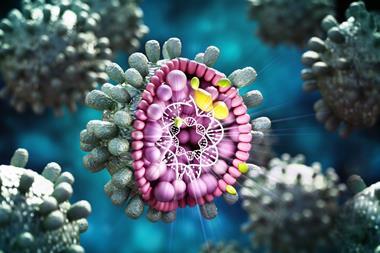
A discovery about Zika’s enzyme, NS2B-NS3, offers promise for therapeutic targets for Zika and other flaviviruses.
Scientists at Sanford Burnham Prebys have found a therapeutic vulnerability of the Zika virus. Viruses have limited genetic material, and few proteins, so all the pieces must work extra hard. The Zika virus is a good example, as it only produces 10 proteins.
The research team discovered that Zika’s enzyme, NS2B-NS3, is a multipurpose tool with two fundamental functions: breaking up proteins (a protease) and dividing its own double-stranded RNA into single strands (a helicase).
Dr Alexey Tersikih, senior author of the paper and Associate Professor at Sanford Burnham Prebys explained: “We found that Zika’s enzyme complex changes function based on how it’s shaped…When in the closed conformation, it acts as a classic protease. But then it cycles between open and super-open conformations, which allows it to grab and then release a single strand of RNA—and these functions are essential for viral replication.”
Zika is an RNA virus belonging to the family of flaviviruses, deadly pathogens including West Nile, dengue fever, yellow fever, and Japanese encephalitis. It is transmitted by mosquitoes and mostly infects uterine and placental cells, making it particularly dangerous for pregnant women. The virus re-engineers the host cells to produce more Zika.
A therapeutic target could be found by understanding Zika on a molecular level. Although it would be challenging to create safe drugs that target the areas of the enzyme needed for protease or helicase functions, as human cells have many similar molecules, a drug that blocks Zika’s conformational changes may be effective. If the complex cannot shape-shift, it cannot perform its essential functions so no new Zika particles would be produced.
Conformational changes
It has long been known that Zika’s essential enzyme is composed of two units: NS2B-NS3pro and NS3hel. NS2B-NS3pro carries out protease functions, cutting long polypeptides into Zika proteins. However, it was only recently discovered how NS2B-NS3pro’s binds single-stranded RNA and aids the separation of the double-stranded RNA during viral replication.
In this study, the researchers used recent crystal structures, protein biochemistry, fluorescence polarisation and computer modelling to analyse NS2B-NS3pro’s life cycle. NS3pro is connected to NS3hel, the helicase, by a short amino acid linker and becomes active when the complex is in its closed conformation. The RNA binding occurs when the complex is open, whereas the complex must transition through the super-open conformation to release RNA.
NS3hel activity drives these conformational changes, as it extends the linker and eventually pulls the NS3pro to release RNA. NS3pro is attached to the inside of the host cell’s endoplasmic reticulum (ER), a key organelle that helps guide cellular proteins to their appropriate destinations via NS2B. Also, while in the closed conformation, it slices the Zika polypeptide, helping generate all viral proteins.
On the other side of the linker, NS3hel separates Zika’s double-stranded RNA and hands a strand over to NS3pro, which has positively charged forks that hold onto the negatively charged RNA.
“There’s a very nice groove of positive charges,” explained Dr Terskikh. “So, RNA just naturally follows that groove. Then the complex shifts to the closed conformation and releases the RNA.”
When NS3hel reaches forward to grab the double-stranded RNA, it pulls the complex with it. However, as the NS3pro is attached to the ER membrane, and the linker can only extend so far, the complex snaps into the super-open conformation and releases RNA. The complex then relaxes back to the open conformation, ready for a new cycle.
When NS3pro detects a viral polypeptide to slice, it forces the complex into the closed conformation, becoming a protease.
As well as providing a potential therapeutic target for Zika, this detailed understanding may be applied to other flaviviruses, which share similar molecular machinery.
The study was published in PLOS Pathogens.