GSK: A Lifetime in Fluorescence
Posted: 4 April 2017 | Carl Haslam, Gabriella Clarke, Luke Greenhough, Peter D Craggs, Ryan P Bingham | No comments yet
GSK’s top investigators explain why the data quality offered by Fluorescence lifetime detection technology for drug discovery applications is superior…

The application and development of new assay technologies plays a critical role in drug identification. In the late 1990s, early drug discovery faced the challenge of ever increasing compound libraries and how best to maximise the impact of the enhanced chemical diversity. To measure the modulation of biological response elicited by individual compounds from these vast collections, many pharmaceutical R&D organisations relied heavily, and still do today, on fluorescent-based in vitro assays, a result of their sensitivity, cost and scalability.
With the implementation of ultra High Throughput Screening (uHTS),1 however, came the additional burden of identifying real biological interventions from those responses driven purely by or masked by compound induced interferences of the fluorescent readout.
In recent years, through the more rigorous classification and QC of compounds, the greater application of red-shifted fluorophores and the implementation of miniaturised fluorescent technologies, or indeed non-fluorescent technologies, focusing on intrinsic molecular properties these issues have only been partly addressed. Confocal fluorescent assay platforms such as Fluorescence Correlation Spectroscopy (FCS)2 and Fluorescence Intensity Distribution Analysis (FIDA)3, focusing on molecular mass and brightness, demonstrated reduced susceptibility to the optical artefacts induced by test compounds compared to other fluorescence-based assay formats4.
These advanced plate-based techniques, as did the non-fluorescent methodologies at that time (such as, Mass Spectrometry and Surface Plasmon Resonance), however, failed to meet the required throughput, due to complex sample handling and/or lengthy read times. Time resolved fluorescence resonance energy transfer (TR-FRET) superseded the aforementioned technologies, in its applicability to uHTS and ability to mitigate effects of compound autofluorescence, making it one of the most preferred assay modalities to date. Nevertheless, the use of specific monoclonal antibodies heightened the biochemical complexity of the assay and more importantly, the cost. Fluorescence lifetime (FLT) measurements offered an equivalent set of advantages to the advanced microplate-based approaches with the scalability required to meet with demands of large compound library screening. Measuring intrinsic lifetime of fluorophores, a property independent of both concentration and volume discrepancies, enables discrimination of the assay response from interfering backgrounds as these, in most instances, decay more rapidly.
Measurement of fluorescent lifetimes has been possible for decades but the availability of truly scalable, 1536 well microplate-based instruments and analysis methodologies had previously limited their application in an uHTS environment.
Measuring lifetime
Early plate readers were based on the two differing approaches to measuring lifetime (i.e Time domain (TD) – Evotec cFLA, Tecan Ultra Evolution and Frequency domain (FD) – LJL Biosystems FLAre) and through collaborations with these vendors, GlaxoSmithKline (GSK) gained access to these technologies with a mind to enhancing screen capability and quality5. Initial experiments demonstrated the power of lifetime measurements to identify true actives rather than optical artefacts and thereby reduce post hit identification cycle times. Early investigations utilising the FLAre, which was based on the intuitive Acquest multi-mode microplate reader, demonstrated an essential requirement for fluorophores with extended lifetimes (τ > 100 ns) due to the phased modulation method of lifetime determination. These novel cryptate-based fluorophores were both difficult to synthesise and to perform target labelling chemistry with compared to the plethora of commonly available fluorophores (Cy5 (τ = 0.8ns), MR121 (τ = 1.2ns), TAMRA (τ = 2.5ns) and Fluorescein (τ = 4ns)).
Autofluorescence
These shorter lifetime fluorophores were, nevertheless, more amenable to the Time Correlated Single Photon Counting (TCSPC) methods of the Evotec and Tecan microplate readers. Conversely their user interfaces were more complex and data output failed to fit the requirements of uHTS; automated deconvolution and recovery to correct optical artefacts was not incorporated. In parallel, although these fluorophores were optimal for single photon counting their utilisation was still sensitive, albeit to a lesser extent, to autofluorescent interferences as a result of longer decaying backgrounds associated with some optically active compounds (lifetime distribution of diverse compound libraries has a long tail extending to >8 ns).
Hence, the broader implementation of fluorescence lifetime outputs for screening over the past two decades has been hampered by the lack of appropriate reporter fluorophores with longer fluorescent lifetimes to those interfering molecules found in compound libraries, a necessity when tackling autofluorescence. This is together with the need for an appropriate uHTS-friendly, bench-top microplate reader, with automated detection and correction of compound interferences.
Almac have developed numerous FLEXYTE FLT reagents that employ a proprietary fluorophore, FLEX17, that has a longer fluorescence lifetime of 17 ns and has been optimised for use with microplate readers from Edinburgh Instruments and more recently TTP Labtech’s Ameon. The Ameon employs real-time decay curve analysis, which allows the detection of thousands of fluorescence decay curves per second that are averaged to allow reporting of highly reproducible and accurate FLT data, without compromising read speeds.
Screening applications
Based on our earlier FLT experiences, GSK have utilised the Ameon microplate reader and FLEXYTE assay reagents to carry out a series of studies to validate their application in miniaturised compound screening applications, including single concentration inhibitor identification and subsequent dose response hit confirmation.
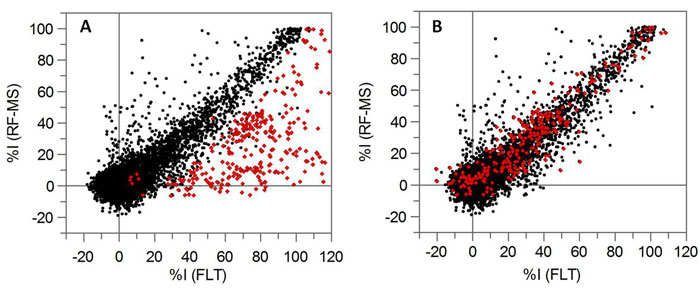

Figure 1: Kinase focused set screening in FLT and RF-MS activity assays. A) Correlation of % inhibition, at a single compound concentration (10 uM), for both FLT and RF-MS activity assays. Red symbols represent those compounds flagged as autofluorescent in the FLT assay. B) as in 1a with the application of correction algorithms to recover the flagged autofluorescent compounds in the FLT data.
As part of this project, we worked extensively with TTP Labtech to exploit the attributes of FLT to develop and implement autofluorescence detection and correction techniques, thereby enabling automatic rescue of real inhibitors that are additionally fluorescent. The correlation in Figure 1A is single concentration screening data generated using two assays to monitor the activity of a tyrosine kinase; one is FLT-based and the other is an Aglient RapidFire, label free, microplate based mass spectrometry approach (RF-MS). There is a good correlation between the data generated using the two assay formats, however, there are also a high number of compounds that are identified as inhibitors using FLT but not using RF-MS (red symbols). Figure 1B is the same single concentration FLT data but the automatic interference and recovery methodology has been applied. What is immediately apparent is that only a limited number of false positive results remain once the rescue algorithms have been applied, affirming FLT, in combination with robust analysis scripts, as a robust fluorescence based readout. 11
The conclusion from these data is that 1 in 8 of the compounds screened in the FLT assay were initially identified as false positives, and using most other fluorescent based techniques would have subsequently been excluded. However, using the interference flag and recovery methodologies embedded in the TTP Labtech Ameon software it was possible to rescue these inhibitors and progress them into further studies.
Conclusion
We have demonstrated through the projects carried out at GSK, utilising the TTP Labtech Ameon microplate reader and Almac FLEXYTE reagents, that there are now viable options to maximise the benefits of FLT and successfully apply this technology in an uHTS environment.
In addition, we have shown that it is possible to identify and rescue fluorescent inhibitor compounds that would have been discarded or missed using other, more established, fluorescent assay technologies. Further studies are ongoing to broaden the application of FLT to a wider diversity of target classes and embed these technologies into GSK’s screening and assay portfolio.


Carl Haslam is the Senior Scientist for Screening, Profiling and Mechanistic Biology UK, GlaxoSmithKline (GSK).


Peter Craggs is an Investigator for Screening, Profiling and Mechanistic Biology UK at GlaxoSmithKline (GSK).


Ryan Bingham is the Group Leader for Screening, Profiling and Mechanistic Biology UK at GlaxoSmithKline (GSK).


Luke Greenhough is a BSc Molecular Biology student at UCL, who is on an industrial experience placement at GlaxoSmithKline (GSK).


Gabriella Clarke is a Bsc Biology undergraduate at Imperial College London, who is on an industrial experience placement at GlaxoSmithKline (GSK).
References
- J. Wölcke and D. Ullmann. Drug Discovery today 2001 (vol 6) 12:637-646.
- K. J. Moore, S. Turconi, S. Ashman, M. Ruediger, U. Haupts, V. Emerick and A. J. Pope. J. Biomol Screening 1999 (Vol.4); 6:335-353.
- U. Haupts, M. Rüdiger, S. Ashman, S. Turconi, R. Bingham, C. Wharton, J. Hutchinson, C. Carey, K. J. Moore and A. J. Pope. J. Biomol Screening 2003 (Vol 8) 1:19-33.
- S. Jäger, N. Garbow, A. Kirsch, H. Preckel, F. U. Gandenberger, K. Herrenknecht, M. Rüdiger, J. P. Hutchinson, R. P. Bingham, F. Ramon, A. Bardera, J. Martin. J. Biomol Screening 2003 (Vol 8) 6:648-659.
- S. Turconi, R. P. Bingham, U. Haupts and A. J. Pope. Drug Discovery today 2001 (vol 6) sup. 1:27-39.
Related topics
Antibodies, Assays, High-Throughput Screening (HTS), Microscopy, Spectroscopy
Related organisations
Almac, Ameon, Edinburgh Instruments, GSK (GlaxoSmithKline)
Related people
Carl Haslam, Gabriella Clarke, Luke Greenhough, Peter D Craggs, Ryan P Bingham