Structure of amyloid fibrils determined with previously unachieved resolution
Posted: 11 September 2017 | Dr Zara Kassam (Drug Target Review) | No comments yet
Researchers have displayed structural details of amyloid structures which may answer many questions on the growth of harmful deposits…
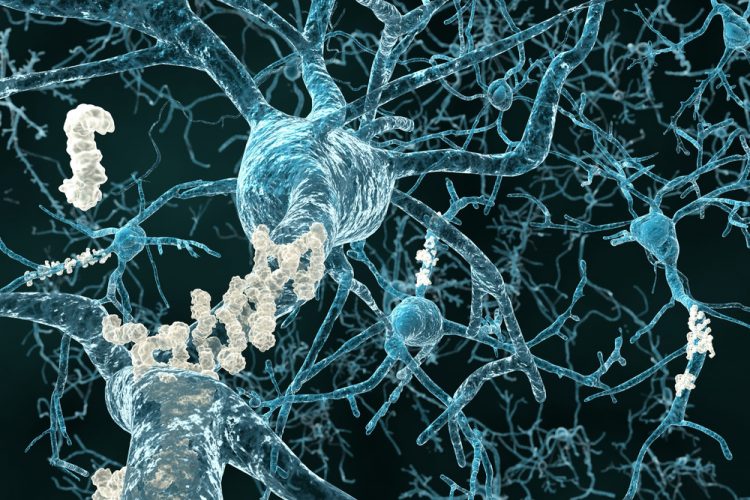

Researchers have displayed previously unknown structural details of amyloid structures which can answer many questions on the growth of harmful deposits and explain the effect of genetic risk factors.
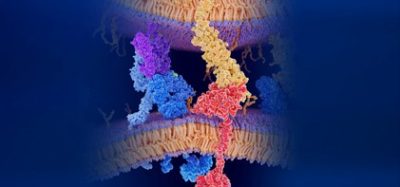
The structure reveals how the many single amyloid beta (Aβ) protein molecules are staggered in layers on top of each other and are arranged into so-called protofilaments. Two of these protofilaments are twinned around each other to form a fibril. If several of these fibrils become entangled, then this gives rise to the typical deposits or plaques that are detected in the brain tissues of Alzheimer’s patients.
“This is a milestone on the road to a fundamental understanding of amyloid structures and the related diseases,” explains Professor Dieter Willbold, director of the Institute of Physical Biology at the Heinrich Heine University Düsseldorf and director of the Institute of Complex Systems (ICS-6) of the Forschungszentrum Jülich.” The fibril structure answers many questions about the mechanism of fibril growth and identifies the role played by a whole series of familial mutations that lead to early onset of Alzheimer’s disease,” he added.
The resolution of 4 angstroms, corresponding to 0.4 nanometres, achieved by the team is within the typical magnitude of atomic radii and atomic bond lengths.
In contrast to previous work, the model shows for the first time the exact position and interactions of the proteins. The Aβ molecules of the entangled protofilaments are thus not at the same level, but like a zipper, they are staggered by half an interval.
Furthermore, the structure elucidates the location and conformation of all 42 amino acid residues of the many individual Aβ protein molecules for the first time.
This novel and detailed structure provides a new basis for understanding the structural effect of many genetic modifications that increase the risk of developing the disease. They stabilize the fibrils – as can now be seen – by changing the blueprint of the protein at defined locations. This e.g. also explains why in nature mice do not develop Alzheimer’s and why a small section of the Icelandic population seems to be more or less resistant to the disease. Their variants of Aβ differ by three or one amino acid residues, respectively, which are apparently important for the stability of the fibrils.
Methodological diversity at the highest technological level
In contrast to the plaques which are typical for the disease discovered by Alois Alzheimer more than 100 years ago, the fibril structure now uncovered cannot be directly observed under the light microscope. It took more than a year to analyse the data the scientists had obtained using the cryo-electron microscopy facility at Maastricht University.
Moreover, measurements using solid-state nuclear magnetic resonance (NMR) spectroscopy and X-ray diffraction experiments helped to supplement and fully support the picture of the fibril structure and validate the data obtained.
“The individual images in cryo-electron microscopy are usually extremely noisy since proteins are very sensitive to electron radiation and the pictures can only be generated with very low radiation intensity,” explains Professor Gunnar Schröder from Forschungszentrum Jülich and Heinrich Heine University Düsseldorf. Using a computer-assisted procedure, he combined thousands of individual images and thus extracted high-resolution structural data from them.
“This is a step that can be very complicated if the sample is heterogeneous, that is to say if it consists of differently formed fibrils. In the past, this was almost always the case with the amyloid fibrils and represented one of the major obstacles for the analysis. However, we now had a fairly unique specimen with very homogeneous fibrils – 90 % of them had the same shape and symmetry,” said Prof Schröder.
Dr Lothar Gremer from and Heinrich Heine University Düsseldorf succeeded in producing the fibril specimen. “The crucial step was to greatly retard the growth of the fibrils in the specimen, from a few hours to several weeks. Thereby the individual Aβ molecules got enough time to arrange themselves into homogeneous fibrils in a very uniform and highly ordered way,” adds Dr Gremer.
Investigations of the fibril specimen by solid-state nuclear magnetic resonance spectroscopy provided additional data to build the model and helped to validate the structure. “NMR enabled us to obtain additional information such as which amino acid residues form salt bridges thus enhancing the stability of the fibrils,” explains Prof Henrike Heise from Heinrich Heine University Düsseldorf and Jülich’s Biomolecular NMR Center. X-ray diffraction experiments supervised by Prof Jörg Labahn at the Centre for Structural Systems Biology in Hamburg additionally confirmed the results.
Related topics
Imaging, Microscopy, Screening
Related conditions
Alzheimer’s disease
Related organisations
Forschungszentrum Jülich, Heinrich Heine University Düsseldorf, Institute of Complex Systems, Institute of Physical Biology
Related people
Dr Lothar Gremer, Professor Dieter Willbold