Trends in contemporary microscopy
Posted: 24 September 2015 | Dimitri Scholz (University College Dublin), Jeremy Simpson (University College Dublin) | No comments yet
In the biomedical arena the imaging of biological samples has long been a mainstay technology, not only furthering our understanding of fundamental biology, but also playing a key role in designing strategies and therapies to combat infection and disease…
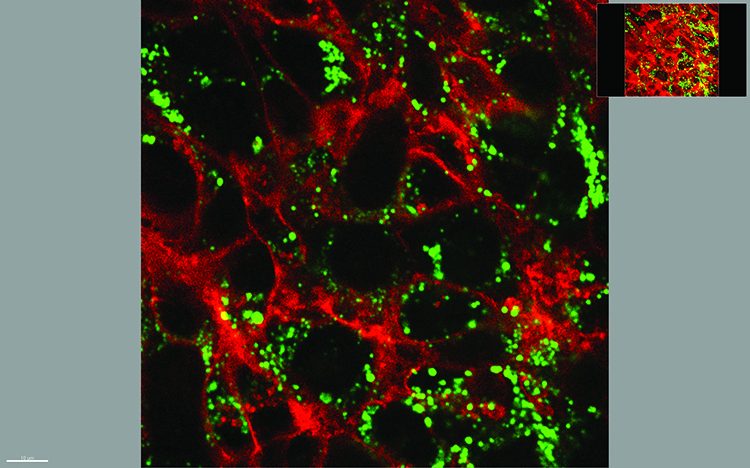
Microscopy is historically two-dimensional, but the world around us is not. We would like to find out more about our samples, with high resolution in all three dimensions, and over time (the 4th dimension) if the sample is living. In the coming years, we expect that three-dimensional microscopy needs will see major technological development. In parallel, improvements in the resolution of non-microscopy 3D imaging approaches, such as ultrasound tomography, micro-CT and positron emission tomography will also undoubtedly occur, thereby narrowing the resolution gap of these technologies with optical microscopy.
In optical microscopy, a light beam passing through a five micrometer-thick specimen (for example tissue or cell layer) can reveal a significant amount of information about the sample; however we cannot yet detect all details and quantify them properly. Although the researcher can learn a lot about the sample by simply scrolling through a number of focal planes, an integrated picture of all dimensions at high (subcellular) resolution cannot easily be captured and interpreted. Instead, several single focal-plane images are often acquired, with the danger that as the thickness of the sample increases, so does the ‘blur’ of each image, as a consequence of the out-of-focus information dominating the image. Calculating the in-focus components from each plane has become possible with motorised focusing and image deconvolution algorithms, but further improvements are still needed here.
Required 3D information can be integrated from multiple physical or optical serial sections. For physical sections, each section needs to be individually collected and separately imaged, and the results compensated for missing sections, areas of compression and unexpected sample rotation, tilting, or twisting. The entire volume is then rendered in silico. Computers with high processing power and specialised software have recently granted us the ability to carry out such tasks, again furthering our capabilities to move towards satisfactory 3D imaging.
A further issue is in determining the best compromise between the analysed volume required and XYZ-resolution. For high XY-resolution of a large area, an XY-scan under high magnification using a motorised stage can be carried out. This no longer represents a technological challenge since in such applications dry objective lenses are usually used. Although oil immersion lenses typically provide higher resolution, conventional mineral oil does not spread evenly around a large sample. Recently, silicon oil and water immersion lenses have become commercially available, thereby reducing the impact of this limitation.

Mrs Tiina O’Neill, Technical Officer, UCD Conway Institute Imaging Core facility, University College Dublin.
However, there are differences in the ability of various software algorithms to accurately ‘stitch’ large numbers of images. Nevertheless, a single scan of a sample sized several cm2 can be completed in a few minutes. The Z-resolution will be defined by the thickness of the section.
The ultimate goal of 3D microscopy – to obtain equal high resolution in each direction – is theoretically achievable; however, this remains a challenging task if the sample volumes are large. For example, in a single experiment in which a 1cm3 tissue sample has to be serially imaged at 500nm steps, 20,000 sections would be required, each of which has to be collected, stained and imaged. Automation is therefore essential. Rather than collecting serial sections, a more promising solution would be block-face imaging combined with serial block shaving. The clear advantage of this approach is the perfect alignment of serial images. The disadvantage lies in the difficulty to label or stain the entire block of tissue. To overcome this, one can either use auto-fluorescence or introduction of a fluorescent marker into the sample prior to its preparation for imaging. In either case, no after-labelling is then required, which is always difficult in a thick sample.
Mouse brain imaging
One of the most successful 3D reconstructions of a reasonably large object (the entire mouse brain) was demonstrated in 20121. The researchers used a combination of fluorescent protein labelling, serial vibratome sectioning and serial two-photon imaging. Vibratomes can section fresh tissue into 50 – 400 µm slices, and two-photon microscopy can image optical planes up to a depth of several hundred micrometers within tissues (Olympus recently announced a lens able to image as deep as 8mm). After vibratome slicing, the block was imaged a number of times at different planes below the surface. Because the Z-resolution of two-photon microscopy is almost as good as the XY resolution, theoretically the entire tissue could be imaged with high resolution of ca. 250-300nm. Practically however, imaging with a two-photon microscope is a time-consuming process. In this ground-breaking example the authors compromised Z-resolution, and still required several days for acquisition of the sample. For fast, relatively low-resolution 3D-reconstructions of larger tissue blocks, single-plain illumination microscopy (SPIM) has a great future. This technique requires no sectioning, but the tissue must be clarified, a technique invented by Werner Spalteholz more than 100 years ago. The clarification of tissue has once again become a popular and helpful tool in 3D-microscopy2.
Researchers have also successfully combined automated cryo-sectioning with block-face fluorescence imaging3. Recently, a commercial system was launched, which combines fluorescence-compatible resin and block-face imaging. The advantage of this approach is the possibility to section as thin as 1µm, and improve Z-resolution correspondingly. Block-face optical (fluorescence) imaging is still in its infancy as a mainstream method of producing 3D image data, but we are likely to see many improvements to this method in the near future, with many of its challenges overcome.
As an alternative to optical imaging, electron microscopy also has the potential to provide 3D image data, and at orders of magnitude of higher resolution than optical techniques. The first commercially-available automated system for 3D scanning electron microscopy (SEM) was Gatan’s 3View® system. The tissue sample is prepared in a similar manner as it would for conventional transmission electron microscopy (TEM); EPON blocks are shaved by an ultramicrotome, but then block-face imaged by a scanning electron microscope. The XY-resolution can be as good as 5nm, Z resolution (section thickness) can be as good as 30µm, and the volume can be as large as 1mm3. In comparison, the FIB-SEM 3D microscopes can reconstruct maximally to 20-50µm in depth, and TEM tomography less than 0.5µm, but with better axial resolution. However, the 3View user has to compromise between the sample volume and the resolution for the sake of time and data size. An alternative approach is the collection of serial sections on conductive slides followed by automated SEM scanning. One device with such functionality presented recently is the ATUMtome. Although the collection of serial sections is likely to be more difficult than serial shaving of blocks, the block-face imaging is technically challenging, and requires extremely high amounts of heavy metals and hard resins.
These technologies have a great future, and they will undoubtedly evolve further towards the reconstruction of ever larger volumes, thereby competing with optical microscopy methods. Current technologies allow resin embedding and automated sectioning of centimeter-scale blocks, which from a medical perspective (for example imaging an entire solid tumour at ultrastructural resolution) is an exciting prospect.
Table-top vs. high-end imaging instrumentation
In the last decade, almost all of the optical and electron microscopy instrumentation manufacturers have brought their cutting-edge devices to the market, and simultaneously offered simpler, cheaper, less flexible but user-friendlier alternatives. This highlights the importance of imaging across the spectrum of researchers, and its need to provide solutions at all price points. In the beginning, these simpler instruments represented only slightly slimmed-down versions of their more mature and sophisticated counterparts, but more recently they have become true ‘all in a box’ table-top devices. One of the most successful examples is the Olympus FV10 confocal microscope. This microscope is simple, robust, user-friendly, and yet it is a real confocal microscope that delivers the same quality of images as his significantly more expensive FV1200 big brother. This table-top microscope also has additional bonuses: it can deliver more images in the same time period, and is not sensitive to ambient light (because it is covered by a box) or vibrations (it has an integrated anti-vibration system). Certain applications, such as photobleaching and fluorescence lifetime imaging, cannot be carried out using this microscope, but its capability covers over 80% of all confocal applications. The microscope requires neither a special room, nor highly trained personnel.

Dr Mattia Bramini, UCD Conway Institute Imaging Core facility, University College Dublin.
Similar developments are occurring in SEM. At the high end, new devices have caught up with the biological TEMs in resolution and have adopted multiple additional detectors for X-rays, photo emission (PEEM) and transmitted electrons (STEM). On the other hand, simple table-top SEMs such as the Hitachi TM3030 grow in popularity. A resolution of ca. 15nm is sufficient for many applications. Priced at only one quarter of the cost of a high-end model, the device covers ca. 80% of biomedical applications, is user-friendly, robust and can be operated much more quickly than the high-end model.
The situation in the TEM market is different. The top-end aberration-corrected microscopes achieve sub-atomic resolution, but also come with multi-million pound price tags. Furthermore, for biomedical applications they are of limited use. Unlike with optical microscopy and SEM, biomedical TEM technologies have not seen significant change in the last 60 years. The introduction of digital cameras into these instruments, which has made our lives easier, has been the most important development, but has not been without its difficulties. Market saturation has been partially caused by the fact that 20-year-old instruments are technologically not significantly different from the latest models. Furthermore, there is now competition from the high-end SEMs that offer similar resolution power (nominally, the middle-range TEMs have a resolution limit of ca. 0.2 nm vs. ca. 1.0-1.5nm for SEMs. However, for biological samples the resolution is limited by sample preparation, and cannot be better than 1nm.). These SEMs can provide similar images in STEM mode and possess additional detectors, for similar or even lower cost.
The first budget-price bench-top TEM arrived in 2012 under the name LVEM5®, from Delong America. Its concept is as simple as revolutionary. A tiny electron gun accelerates the electrons to only 5Kev, which reduces many safety issues. The low-magnification image is projected on to a small fluorescent screen, on top of which a fluorescence light microscope with a 40x dry lens is mounted. The entire microscope system is no larger than a research-grade upright optical microscope and with a list price that is comparable. Additional advantages include an SEM detector and fewer requirements for contrasting with heavy metals because of its lower electron acceleration power. The contrast will be achieved by light biological elements, which might bring us new valuable information.
Automated imaging
Another area of rapid change in imaging is its automation, particularly with regard to generating enormous data sets for systems biology. This technology is generally referred to as high-content screening (HCS) microscopy, a term introduced by Cellomics, the developers of the first commercial automated microscopes. In the last two decades all of the traditional optical microscope manufacturers have also developed their own HCS systems, including both basic user-friendly models and high-end high-functionality variants. All such systems share the ability to be able to automatically acquire images, with their automation components offering ever faster multi-well plate loading, stage movement, focusing, filter changing and image acquisition. Such instruments can capture thousands of images in a few hours, and have therefore been extremely useful for basic researchers to exploit information from the human genome, and also for pharmaceutical companies wishing to test huge compound libraries for particular effects on cells. This imaging technology is now playing a major role in the drug discovery and development pipeline, not least in helping identify and test novel therapeutic carriers, such as nanoparticles, prior to in vivo testing4.
However, the latest models are now beginning to add advanced functionalities, going beyond image acquisition, formerly only possible on high-end manually operated confocal microscopes. For example, the Opera Phenix system from Perkin Elmer, a fully automated spinning disk confocal microscope, can be configured with hardware and software to allow it to carry out Förster resonance energy transfer (FRET) experiments in an automated manner. This advanced modality can be used to quantitatively measure protein-protein interactions in living cells, making this technique now theoretically possible on a proteome-wide scale, thereby ultimately helping to reveal how all proteins interact in the cellular environment. HCS is also expected to be an imaging technology that transitions from 2D to 3D. Similar to what we have described above, there are technological challenges to overcome, but as 3D cellular models, such as cell cysts and spheroids, become more commonplace in biological research, it is inevitable that HCS approaches will adapt to be able to extract single cell information from these potentially useful tissue models.
Regardless of the imaging modality employed, all recent advancements have had one outcome, namely that the volume of image data produced per ‘single experiment’ has grown exponentially. Inevitably this means that the modern researcher needs to rely ever more on automated image analysis, rather than slow manual inspection of images. HCS has led in this regard, with software tools and analysis algorithms becoming increasingly accurate at identifying individual cells and even organelles. As the volumes of image data grow, particularly from 3D data sets, these software tools will need to keep pace if we are ever to make sense of all the images that we create, and the important information that they contain.
Biographies
 DR. DIMITRI SCHOLZ is Director of Biological Imaging at the Conway Institute, UCD. He introduced advanced methods in light and electron microscopy and in image analysis and is involved in more than 200 projects and teaching. From 2003 to 2088 he was Research Assistant Professor at the Medical University of South Carolina (MUSC) in Charleston (US) and Principle Investigator, examining intracellular transport of mRNA. Prior to this, between 1992 and 2003 he was Senior Scientist at Max-Planck Institute in Bad Nauheim, Germany, where he studied an animal model of peripheral angiogenesis and undertook clinical research. He has authored 54 peer-reviewed publications.
DR. DIMITRI SCHOLZ is Director of Biological Imaging at the Conway Institute, UCD. He introduced advanced methods in light and electron microscopy and in image analysis and is involved in more than 200 projects and teaching. From 2003 to 2088 he was Research Assistant Professor at the Medical University of South Carolina (MUSC) in Charleston (US) and Principle Investigator, examining intracellular transport of mRNA. Prior to this, between 1992 and 2003 he was Senior Scientist at Max-Planck Institute in Bad Nauheim, Germany, where he studied an animal model of peripheral angiogenesis and undertook clinical research. He has authored 54 peer-reviewed publications.
 JEREMY SIMPSON carried out his PhD work at the University of Warwick (UK). After post-doctoral work at the Scripps Research Institute in San Diego (USA) and the ICRF in London (UK), he was awarded a long term EMBO fellowship and spent nine years at the European Molecular Biology Laboratory (EMBL) in Heidelberg (Germany). In 2008 he was appointed as Professor of Cell Biology at University College Dublin, Ireland. His lab currently applies high-throughput imaging technologies to study trafficking between the ER and Golgi complex, the internalisation pathways taken by synthetic nanoparticles on exposure to cells, and how the cytoskeleton responds to cellular exposure to nanomaterials and nanosurfaces. His lab also develops novel software tools for image analysis. He has authored over 80 peer-reviewed articles, including articles in Nature Cell Biology and Nature Methods, and a number of book chapters. He runs the UCD Cell Screening Laboratory (www.ucd.ie/hcs) and he is currently is the Head of School of Biology & Environmental Science and the Vice Principal (International) in the UCD College of Science.
JEREMY SIMPSON carried out his PhD work at the University of Warwick (UK). After post-doctoral work at the Scripps Research Institute in San Diego (USA) and the ICRF in London (UK), he was awarded a long term EMBO fellowship and spent nine years at the European Molecular Biology Laboratory (EMBL) in Heidelberg (Germany). In 2008 he was appointed as Professor of Cell Biology at University College Dublin, Ireland. His lab currently applies high-throughput imaging technologies to study trafficking between the ER and Golgi complex, the internalisation pathways taken by synthetic nanoparticles on exposure to cells, and how the cytoskeleton responds to cellular exposure to nanomaterials and nanosurfaces. His lab also develops novel software tools for image analysis. He has authored over 80 peer-reviewed articles, including articles in Nature Cell Biology and Nature Methods, and a number of book chapters. He runs the UCD Cell Screening Laboratory (www.ucd.ie/hcs) and he is currently is the Head of School of Biology & Environmental Science and the Vice Principal (International) in the UCD College of Science.
References
- Ragan T, Kadiri LR, Venkataraju KU, Bahlmann K, Sutin J, Taranda J, Arganda-Carreras I, Kim Y, Seung HS & Osten P. Serial two-photon tomography: an automated method for ex-vivo mouse brain imaging. Nat Methods. 2012; 9, 255-258
- Tomer R, Ye L, Hsueh B. & Deisseroth K. Advanced CLARITY for rapid and high-resolution imaging of intact tissues. Nature Protocols. 2014; 9, 1682–1697
- Roy D, Steyer G, Gargesha, M, Stone, M & Wilson D. 3D Cryo-Imaging: A Very High-Resolution View of the Whole Mouse. Anat Rec. 2009; 292, 342–351
- Brayden DJ, Cryan SA, Dawson KA, O’Brien, PJ & Simpson JC. High-content analysis for drug delivery and nanoparticle applications. Drug Discov. Today. 2015; doi: 10.1016/j.drudis.2015.04.001
Related topics
Imaging, Microscopy, Positron emission tomography (PET), Technology