Bringing native analysis into the fast-moving world of biologics
Posted: 12 June 2019 | Christopher Nortcliffe - SCIEX | No comments yet
For the past thirty years, native mass spectrometry has grown in both scope and reach in labs across the globe to encompass larger and more challenging molecular complexes. However, up until now, the throughput of these techniques has been slow and manual. Christopher Nortcliffe discusses ways that native analysis is advancing the field of mass spectrometry.
The integration between column and instrument manufacturers seeks to unlock new rapid native analysis, bringing the technology firmly into the hands of the pharmaceutical industry. The analysis of proteins in the gas phase underwent an expansion of Cambrian magnitude with the development of soft ionisation techniques such as electrospray ionisation (ESI) and matrix-assisted laser desorption ionisation (MALDI).1-5 These techniques allowed large fragile biomolecules to be transmitted into the gas phase along with the retention of some protein-protein and protein-ligand interactions. However, despite the ‘softness’ of these techniques, most of the work carried out still relied upon the use of denaturing solvents such as acetonitrile and methanol. Therefore, although the biomolecules entered the gas phase intact, all their structural and stoichiometric data had been lost.
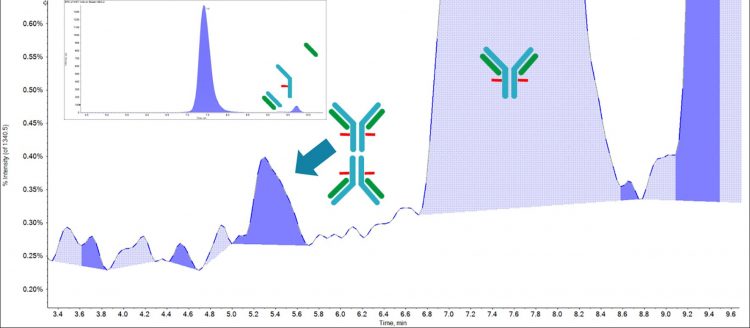
Figure 1: Separation of the NIST mAb with distinct peaks for dimer, monomer and fragment species
Development of new volatile buffer techniques and improvements in instrumentation have enabled the transmission of proteins and non-covalent complexes into the gas phase whilst retaining their solution phase structure. This technique was coined native or ‘native-like’ mass spectrometry.6 However, there was debate over the viability of the data from these techniques when compared with more established techniques such as NMR, X-ray crystallography and electron microscopy. Combining native mass spectrometry with fragmentation techniques such as collision-induced dissociation and ion mobility allowed it to find its niche. This enabled these complexes to be analysed far more rapidly than similar structural techniques and could capture dynamic interactions taking place on time scales that were previously inaccessible.7
Numerous key developments in the field of native mass spectrometry have developed into new areas of
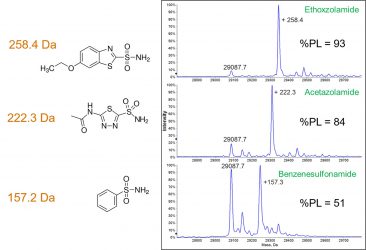
Figure 2: Reconstructed
mass spectrum of carbonic anhydrase with the addition of three ligand species. Degree of binding was determined via peak area of bound versus unbound species.
structural biology and have answered fresh questions. Fragmentation techniques have developed such as electron capture dissociation (ECD) and electron transfer dissociation (ETD), which allow extensive coverage of biomolecules during fragmentation. These have been particularly useful when employed with protein cross-linking studies. A recent addition to this field of surface-induced dissociation pioneered by the Wyzocki group has paved the way for subunit stoichiometries and folding to be interrogated to exciting new frontiers.8 Other improvements have arisen since instrumentation with ion-mobility has become commercially available from several vendors in various orientations, including differential, drift time and travelling wave. These hybrid techniques give an orthogonal degree of separation to mass spectrometry for the separation of complex samples. Modified instrumentation specifically tuned to transmit large complexes into the gas phase has allowed immense macromolecular structures to be studied including whole intact viral capsids.9
Despite progress being made in the field, most of the work has taken place in academic institutions and the technique hasn’t really migrated into the pharmaceutical industry, despite the abundance of mass spectrometry technology already in use. A key reason for this is the individual way most native experiments have been performed on very low flow techniques. A common way to undertake a native experiment is by filling a pulled glass capillary with the sample before applying a voltage differential. These tips must be filled individually by hand, are fragile and prone to blockages caused by aggregates or insufficient sample preparation. Steps have been taken to give this process higher throughput, for example the development of the Advion Nanomate.10 However, this requires a specific piece of instrumentation to be set up for native analysis and can be expensive to run in a high-throughput manner.
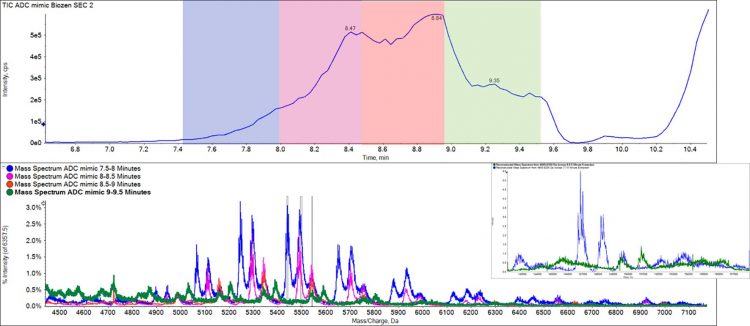
Figure 3: Chromatogram of an ADC mimic showing four distinct elution zones. Mass spectrum extracted from each elution zone shows the separation of different DAR isoforms based upon size.
The pharmaceutical industry’s reliance on high-throughput batches and LC technology means the benefits of native analysis have so far been outweighed by operational logistics. However, there is now pressure for samples to be analysed in a native way with the development of new biotherapeutics. Three major areas are: new antibody drug conjugates with cysteine linked payloads; increased desire to understand and control protein aggregation; and the analysis of the specific interactions between small molecule drugs and protein targets. These burgeoning areas rely upon non-covalent interactions, which cannot be maintained through traditional LC-MS approaches. Potential solutions are being developed by the union of mass spectrometry and chromatography. The key factors driving this are:
- High-throughput methods that fit within the already established pharmaceutical pipeline
- Methods on routine instrumentation without need for modification or specific add-ons
- Robustness and reliability leading to confidence in results.
Size exclusion chromatography (SEC) is not a new technique and has been utilised for many years in the biopharma industry with UV detection. SEC separates molecules based upon their size with large molecules unable to enter the pores of the column and therefore experiencing a shorter path length than smaller molecules. Due to this, any non-volatile salts present in the sample buffer should be held on the column to be eluted to waste later, whilst the biological sample elutes off the column earlier. Larger protein aggregates can also be separated from monomers.
Figure 1 shows the example of aggregate separation utilising the BioZen Sec 2 column. There is a clear peak eluting around 5.3 minutes, which shows only signal from the protein dimer with no monomer eluting. There is then a major peak around 7.5 minutes with evidence of monomer followed by two distinct fragment species. The m/z of each of these species can then be confirmed using the mass spectrometer.
Figure 2 outlines a series of experiments utilising carbonic anhydrase along with a series of known carbonic anhydrase inhibitors. Not only are the protein-ligand interactions maintained into the gas phase, but the stoichiometry of binding can be used to determine a binding strength.
Figure 3 and Figure 4 show work on an ADC mimic obtained from Sigma Aldrich, which has a fluorescent molecule bound to cysteine residues. Under standard reverse phase conditions, the heavy and light chains of the antibody separate, and the overall DAR ratio can’t be observed. Under native conditions the complex is kept intact and the separation of various DAR forms through the SEC shows the different elution profiles of each form, contributing to the understanding of ADC formation.
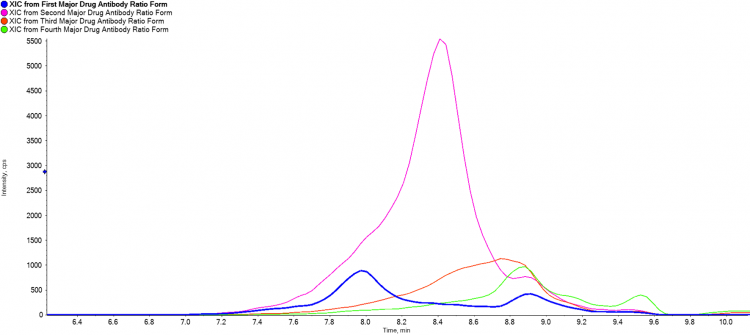
Figure 4:Extracted ion FIGURE 4 chromatogram of four DAR isoforms across a single charge state showing differences in elution profile for BioZen SEC 2 column.
In conclusion, this new workflow seeks to address issues with certain biomolecules and lowers the hurdles associated with the adaption of native mass spectrometry.
Biography
CHRISTOPHER NORTCLIFFE completed his undergraduate master’s at Edinburgh University before joining a Doctoral training in Proteomics and Cell Technology. During this time, he focused on analytical techniques with an emphasis on mass spectrometry using QToFS and FT-ICR. He conducted his PhD under Professor Perdita Barran at Manchester University, titled “Analysis of Non-Covalent Complexes using Mass Spectrometry Techniques”. This focused on the application of ion mobility-MS. Upon finishing his studies, he started working at SCIEX in 2016 as an Application Specialist in Biologics and Biopharma.
References
- Shelimov KB, Clemmer DE, Hudgins RR, Jarrold MF. J. Am. Chem. Soc. 1997; 119: 2240–2248
- Badman ER, Hoaglund-Hyzer CS, Clemmer DE. Anal. Chem. 2001; 73: 6000–6007
- Valentine SJ, Clemmer DE. J. Am. Soc. Mass Spectrom. 2002; 13: 506–517,
- Gidden J, Ferzoco A, Baker ES and Bowers MT. J. Am. Chem. Soc. 2004: 126; 15132–15140,
- Bernstein SL, Liu DF, Wyttenbach T, Bowers MT, Lee JC, Gray HB and Winkler JR. J. Am. Soc. Mass Spectrom. 2004: 15; 1435–1443.
- van den Heuvel RH, Heck AJ,. Curr. Opin. Chem. Biol. 2004: 8; 519 –526.
- Kaddis CS, Loo JA. Anal. Chem. 2007: 79; 1778 –1784.
- Wysocki VH, Joyce KE, Jones CM, Beardsley RL. J Am Soc Mass Spectrom. 2007: 19(2); 190–208.
- Snijder J, Rose RJ, Veesler D, Johnson JE, Heck AJ. Angewandte Chemie 2013: 52(14); 4020-4023.
- Constans A, 2003 The Scientist, 17(7), 49-50.
Related topics
Analytical Techniques, Biologics, Biotherapeutics, Mass Spectrometry, Molecular Biology
Related organisations
SCIEX