Biomarkers that could give better cancer survival estimates
Posted: 9 June 2016 | Victoria White, Digital Content Producer | No comments yet
A new method, developed by UCLA scientists, could eventually lead to a way to improve accuracy of how well a patient will respond to treatment…
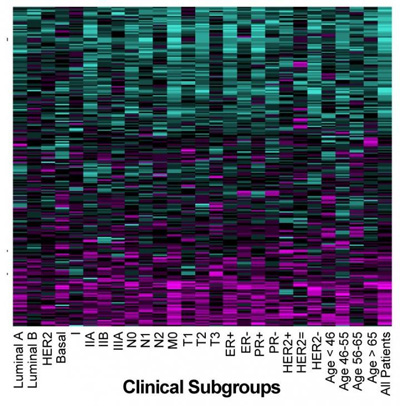
SURVIV analysis of breast cancer isoforms. Blue isoforms are associated with longer survival times, and magenta isoforms with shorter survival times. CREDIT: Yi Xing
Researchers from UCLA have used data about patients’ genetic sequences to produce more reliable projections for survival time and how they might respond to possible treatments.
The technique is an innovative way of using biomedical big data to achieve precision medicine – giving doctors the ability to better tailor their care for each individual patient.
The approach is likely to enable doctors to give more accurate predictions for people with many types of cancers. In this research, the UCLA scientists studied cancers of the breast, brain (glioblastoma multiforme), lung, ovary and kidney.
The new method analyses various gene isoforms using data from RNA molecules in cancer specimens. This RNA sequencing (RNA-seq) reveals the presence and quantity of RNA molecules in a biological sample. In the method developed at UCLA, scientists analysed the ratios of slightly different genetic sequences within the isoforms, enabling them to detect important but subtle differences in the genetic sequences. In contrast, the conventional analysis aggregates all of the isoforms together, meaning that the technique misses important differences within the isoforms.
SURVIV (for “survival analysis of mRNA isoform variation”) is the first statistical method for conducting survival analysis on isoforms using RNA-seq data, said Yi Xing, a UCLA associate professor of microbiology, immunology and molecular genetics.
200 isoforms associated with survival time
The researchers report having identified some 200 isoforms that are associated with survival time for people with breast cancer; some predict longer survival times, others are linked to shorter times. Armed with that knowledge, scientists might eventually be able to target the isoforms associated with shorter survival times in order to suppress them and fight disease, Xing said.
The researchers evaluated the performance of survival predictors using a metric called C-index and found that across the six different types of cancer they analysed, their isoform-based predictions performed consistently better than the conventional gene-based predictions.
The result was surprising because it suggests that isoform ratios provide a more robust molecular signature of cancer patients than overall gene abundance, said Xing, director of UCLA’s bioinformatics doctoral programme and a member of the UCLA Institute for Quantitative and Computational Biosciences.
“Our finding suggests that isoform ratios provide a more robust molecular signature of cancer patients in large-scale RNA-seq datasets,” he said.
The researchers studied tissues from 2,684 people with cancer, and they spent more than two years developing the algorithm for SURVIV.
According to Xing, a human gene typically produces seven to 10 isoforms. “In cancer, sometimes a single gene produces two isoforms, one of which promotes metastasis and one of which represses metastasis,” he said, adding that understanding the differences between the two is extremely important in combatting cancer.
Related topics
Big Data, Biomarkers, Genomics, Oncology
Related conditions
Glioblastoma multiforme
Related organisations
UCLA