Researchers link SARS-CoV-2 PLpro to COVID-19 symptoms
Posted: 10 June 2021 | Victoria Rees (Drug Target Review) | No comments yet
Researchers have shown that the SARS-CoV-2 PLpro cuts human proteins so could potentially cause a range of COVID-19 symptoms.
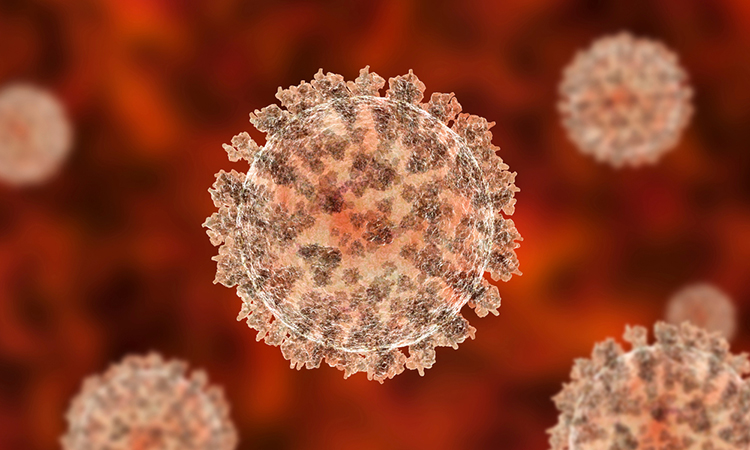

Researchers have found that the SARS-CoV-2 papain-like protease (PLpro) has a range of targets in the human proteome, including proteins involved in cardiovascular function, blood clotting and inflammation. According to the team from the US Naval Research Laboratory, the findings suggest a link between the inactivation of these proteins and COVID-19 symptoms.
The researchers explain that PLpro plays an essential role in processing viral proteins needed for SARS-CoV-2 replication. In addition, the enzyme can cut and inactivate some human proteins important for an immune response.
Viruses like SARS-CoV-2 make multiple proteins as one long “polyprotein”. Proteases recognise specific amino acid sequences in this polyprotein and cut them to release individual proteins. However, some human proteins also contain these sequences (known as homologous host-pathogen sequences [SSHHPS]), including ones involved in generating the innate immune response, which could help protect the virus from the host. The researchers wanted to comprehensively identify human proteins that contain SSHHPS, examine their functions and see whether PLpro can cleave them in a test tube.
The researchers developed a computational method to search a database of all known human proteins for sequences similar or identical to the SARS-CoV-2 SSHHPS. The analysis revealed that the proteins with highest sequence identity were those that had cardiovascular, inflammatory, kidney, respiratory or blood-related functions. For example, two of the proteins containing SSHHPS were cardiac myosins, one was an anti-coagulant and another was an anti-inflammatory protein. Inactivation of these proteins by PLpro is consistent with COVID-19 symptoms of heart damage, blood clots and inflammation. The team confirmed that PLpro could cut these protein sequences in vitro.
Performing the same analysis on SSHHPS for the Zika viral protease identified proteins associated with neurological development and disorders, consistent with Zika symptoms. The team say these results suggest that the symptoms and virulence of viruses can be predicted directly from their genomic sequences.
The study is published in ACS Infectious Diseases.
Related topics
Disease Research, Molecular Targets, Protein, Proteomics
Related conditions
Covid-19
Related organisations
US Naval Research Laboratory