Next generation sequencing for cancer precision medicine
Posted: 17 September 2019 | Aino Palva (University of Helsinki), Lauri Paasonen (UPM Biomedicals), Pekka Ellonen (Helsinki Institute for Life Sciences), Piia Mikkonen (Institute for Molecular Medicine Finland), Vilja Pietiäinen (Institute for Molecular Medicine Finland) | No comments yet
The systematic successful treatment of cancer still eludes us and in an effort to refine this area of targeted medicine, Lauri Paasonen and colleagues explore the potential of using patient-derived cells (PDCs) for devising a personalised treatment strategy for solid tumours.


There are many successful cancer therapies on the market but still, depending on the cancer subtype and stage of the disease, many patients do not fully respond to treatment or can later present with a drug-resistant metastatic disease.
However, many new targeted therapies have been developed but their effectiveness for different cancer types has not been evaluated. In the precision medicine (PM) approach, a vast amount of clinical and omics data is collected from a patient in order to find potential drug targets and to translate the results back to the clinic. Importantly, this research may aid the understanding of drug resistance arising from inter/intratumour heterogeneity. While the genomic alterations, such as somatic driver mutations, have been thoroughly investigated, only a few can be targeted with drugs. This has led to the development of the functional PM approach, which has been successful, for example, in leukaemias. Here, the patient-derived cancer cells (PDCs) can be directly used – without the need for cell culture steps – for ex vivo drug sensitivity and resistance testing.1-3 The response of PDCs is tested against hundreds of approved and investigational drugs, depending on the platform. The combination of the drug response data with phenotypic and genotypic omics data and clinical information about the patient enables a personalised treatment strategy to be developed.


Biomarkers aren’t just supporting drug discovery – they’re driving it
FREE market report
From smarter trials to faster insights, this report unpacks the science, strategy and real-world impact behind the next generation of precision therapies.
What you’ll unlock:
- How biomarkers are guiding dose selection and early efficacy decisions in complex trials
- Why multi-omics, liquid biopsy and digital tools are redefining the discovery process
- What makes lab data regulatory-ready and why alignment matters from day one
Explore how biomarkers are shaping early drug development
Access the full report – it’s free!
NGS in precision medicine
Ex vivo drug testing with genetically- and phenotypically-characterised PDCs from solid tumours allows us to tailor patient-specific treatments”
Next generation sequencing (NGS) has had a big impact on genomic research. High-throughput sequencing technologies have emerged that fulfil the need for sequencing of large gene quantities, such as the whole exome with ~23,000 different genes, and for the quantitative analysis of mutation frequencies. This has also enabled genome sequencing to be used as a clinical tool.7 The characterisation of somatic mutations and copy number variations in PDCs means they can be properly matched with the tumour tissue they are derived from and is a crucial step for any further PDC-based assay.8
Whole exome sequencing or targeted cancer panel sequencing can be performed for PDCs derived both from liquid biopsies and solid tumour samples, but the limited quantity of PDCs often sets certain requirements for the assays. In addition, the isolation of DNA of good quality and adequate quantity from solid tumour PDCs cultured in a 3D matrix can be challenging, as the matrix used may not be easy to remove or can cause steric hindrance during the isolation. Typically, the DNA isolation is performed using commercial column extraction methods, but when the cells are cultured in 3D, the properties of 3D matrices may not allow the proper filtration of samples through the DNA isolation column, resulting in an insufficient yield and poor DNA quality. However, by choosing a suitable 3D culture matrix and protocol, the isolation of DNA for NGS library construction can be performed.
DNA isolation and NGS of renal cancer
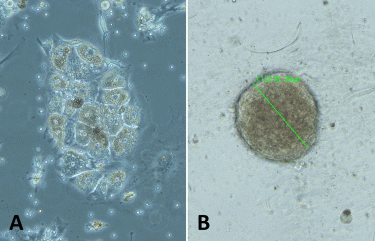

Figure 1: The example images of renal cancer PDCs grown A) in 2D as a monolayer, and B) in 3D in GrowDex after cellulase enzyme treatment (spheroid diameter 216μm). Samples were imaged with Leica S80 microscope, 10x objective.
PDCs cultured in 3D in GrowDex We have optimised the growth of renal PDCs in 2D as a monolayer and in 3D using different matrices (Figure 1). These matrices include GrowDex, a plant-based cellulose hydrogel that has recently been demonstrated as a suitable matrix for the 3D culturing of various cell types.9,10 It is an animal-free, clean and tunable hydrogel, which can be degraded using cellulase enzyme without impacting the human cells.9 As matrices used to support the growth of PDCs in 3D may hinder the typical column-based DNA purification prior to sequencing, we streamlined the workflow for whole exome sequencing (WES) from a low amount of renal cancer PDCs grown as spheroids in GrowDex. As a control, NGS was also performed for the cells cultured in a traditional 2D monolayer setup. The detailed protocol can be found as a separate application note.11
Shortly after, renal tumour tissue samples were dissociated and cultured in 2D according to previously published protocols12,13 or in 0.4 percent GrowDex hydrogel in complete growth medium in 24- or 96-well plates. GrowDex was degraded with cellulase enzyme GrowDase and DNA was isolated with Dynabeads DNA Direct kit. 150ng of DNA was used for library construction and exome sequencing was performed using Roche HyperCap MedExome. Resulting libraries were sequenced with Illumina Hiseq2500 Rapid PE100 runs. Sequence analysis and variant calling was performed using in‑house tools.14
We observed that removal of the 3D matrix is required for an efficient isolation of DNA from 3D-cultured cells by Dynabeads isolation kit. GrowDase enzyme treatment enabled the effective release of the PDC spheroids from GrowDex (Figure 1B) and the robust processing of the spheroids combined with Dynabeads on the magnetic rack. The success of exome capture is critically dependent on the amount and quality of input DNA. In general, a good yield of DNA was obtained from the PDCs cultured 3D in GrowDex; sufficient for WES or any other type of NGS assay. With the DNA isolated from 3D cultures, we were able to generate WES data with somatic resolution (mean target coverage >80x).11 The comparison of sequencing results at the chromosomal level with the Integrative Genomics Viewer15,16 revealed similar coverage in the sequencing of DNA isolated from PDCs cultured either traditionally in 2D monolayer on the plastic or 3D in GrowDex (Figure 2).
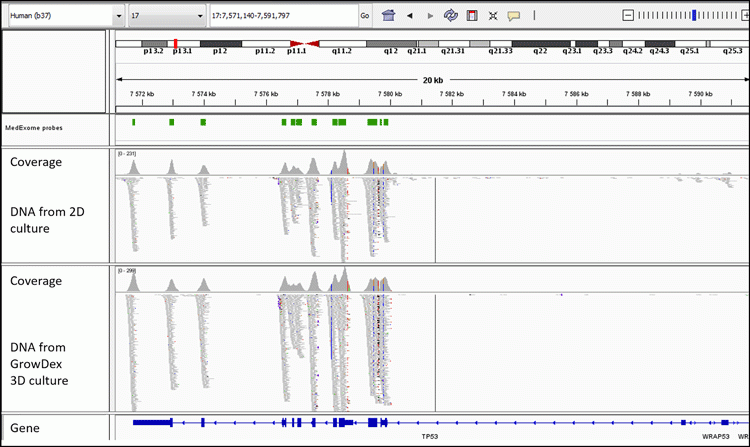

Figure 2: Integrative Genomics Viewer image shows the comparable sequencing coverage of TP53 for DNA isolated from 2D- and 3D-cultured PDCs.11
Summary and conclusions
Novel 3D-culturing methods for PDCs enable more physiologically-relevant ex vivo cancer models. Fast and cost-effective genome sequencing of PDCs is needed for characterisation of the cells and the clinical implementation of ex vivo drug efficacy screening results in PM. Here, the renal cancer PDCs cultured in GrowDex in 3D were made accessible for lysis and bead-based DNA recovery with cellulase enzyme treatment degrading the nanofibrillar cellulose network. Additional purification steps typically present in DNA isolation were needed and the recovered amount and quality of DNA was sufficient for NGS library construction and exome sequencing. In conclusion, the development of fast, straightforward assay protocols optimised for 3D PDC spheroids/ organoids can further improve their utilisation for PM, drug discovery and other applications.
About the authors




Piia Mikkonen, Institute for Molecular Medicine Finland (FIMM), Pekka Ellonen, Helsinki Institute for Life Sciences (HiLIFE) and Aino Palva, University of Helsinki, Finland were also additional authors.
Acknowledgments:
The authors would like to thank FIMM Genomics Core Unit (HiLIFE, University of Helsinki), funded by BioCenter Finland. The tissue samples were obtained through DEDUCER study (the Development of diagnostics and treatment of urological cancers; main investigator in the clinic: Associated Professor A Rannikko, University of Helsinki; with the approved study permissions for HUS/71/2017, 26.04.2017, ethical committee approval 15.03.2017 Dnro 154/13/03/02/2016, and patient consents). This work is based on research collaboration between academy and UPM, supported by UPM-Kymmene Corporation, Finland.
References:
- Pemovska T, Kontro M, Yadav B, Edgren H, Eldfors S, Szwajda A, Almusa H. Individualized Systems Medicine (ISM) strategy to tailor treatments for patients with chemorefractory acute myeloid leukemia. Cancer Discov. 2013 Dec;3(12):1416-29.
- Pemovska T, Johnson E, Kontro M, Repasky GA, Chen J, Wells P, Cronin CN, McTigue M, Kallioniemi O, Porkka K, Murray BW, Wennerberg K. Axitinib effectively inhibits BCR-ABL1 (T315I) with a unique binding conformation. Nature 2015, 519(7541):102-5.
- Snijder B, Vladimer GI, Krall N, Miura K, Schmolke AS, Kornauth C, Lopez de la Fuente O, Choi HS, van der Kouwe E, Gültekin S, Kazianka L, Bigenzahn JW, et al. Image based ex-vivo drug screening for patients with aggressive haematological malignancies: interim results from a single-arm, open-label, pilot study. Lancet Haematol. 2017 Dec;4(12).
- Drost J and Clevers H. Organoids in cancer research. Nat Rev Cancer. 2018 Jul;18(7):407-418.
- Horvath P, Aulner N, Bickle M, Davies A, Del Nery E, Ebner D, Montoya MC, Östling P, Pietiäinen V, Price L, Shorte SL, Turcatti G von Schantz-Fant C, Carragher NO, the EuCAI (European Cell Based Assay Interest Group). Screening out irrelevant cell-based models of disease, Nat Rev Drug Discov. 2016 Nov;15(11):751-769.
- Tuveson D, Clevers H. Cancer modeling meets human organoid technology. Science. 2019 Jun 7;364(6444): 952-955.
- Goodwin S, McPherson JD, McCombie WR. Coming of age: ten years of next-generation sequencing technologies. Nature Reviews Genetics 2016 17, pages 333–351.
- Friedman AA, Letai A, Fisher DE, Flaherty KT. Precision medicine for cancer with next-generation functional diagnostics. Nature Reviews Cancer 2015 volume15, pages 747–756.
- Lou Y-R, Kanninen L, Kuisma T, Niklander J, Noon LA, Burks D, Urtti A, Yliperttula M. The Use of Nanofibrillar Cellulose Hydrogel as a Flexible Three-Dimensional Model to Culture Human Pluripotent Stem Cells. Stem Cells and Development, 2014, Volume 23, Number 4.
- Rinner B, Gandolfi G, Meditz K, Frisch M, Wagner K, Ciarrocchi A, Torricelli F, Koivuniemi R, Niklander J, Liegl-Atzwanger B, Lohberger B, Heitzer E, Ghaffari-Tabrizi-Wizsy N, Zweytick D, Zalaudek I. MUG-Mel2, a novel highly pigmented and well characterized NRAS mutated human melanoma cell line. Scientific Reports 2017 7, 2098.
- Mikkonen P, Palva A, Ellonen P, Pietiäinen V. NGS compatible streamlined DNA recovery from 3D cell cultures in GrowDex. https://www.upmbiomedicals.com/resourcecenter/application-notes/ngs-compatible-streamlined-dnarecovery-from-3d-cell-cultures-in-growdex/ 2018.
- Saeed K, Rahkama V, Eldfors S, Bychkov D, Mpindi JP, Yadav B, Paavolainen L, Aittokallio T, Heckman C, Wennerberg K, Peehl DM, Horvath P, Mirtti T, Rannikko A, Kallioniemi O, Östling P, Af Hällström TM. Comprehensive Drug Testing of Patient-derived Conditionally Reprogrammed Cells from Castration-resistant Prostate Cancer. Eur Urol. 2017 Mar;71(3):319-327.
- Saeed K, Ojamies P, Pellinen T, Eldfors S, Turkki R, Lundin J, Järvinen P, Nisen H, Taari K, Af Hällström TM, Rannikko A, Mirtti T, Kallioniemi O, Östling P. Clonal heterogeneity influences drug responsiveness in renal cancer assessed by ex vivo drug testing of multiple patient-derived cancer cells. Int J Cancer 2019 Mar;144(6), pp. 1356-1366.
- Sulonen AM, Ellonen P, Almusa H, Lepistö M, Eldfors S, Hannula S, Miettinen T, Tyynismaa H, Salo P, Heckman C, Joensuu H, Raivio T, Suomalainen A, Saarela J. Comparison of solution-based exome capture methods for next generation sequencing. Genome Biology 2011 Sep 28;12(9):R94.
- Robinson JT, Thorvaldsdóttir H, Winckler W, Guttman M, Lander ES, Getz G, Mesirov JP. Integrative Genomics Viewer. Nature Biotechnology 2011 29, 24–26.
- Thorvaldsdóttir H, Robinson JT, Mesirov JP. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Briefings in Bioinformatics 2013 14, 178-192.
Related topics
Drug Targets, Next-Generation Sequencing (NGS), Oncology, Precision Medicine, Sequencing, Targets