Proteomics: cracking the molecular mystery of diseases and therapeutics
Posted: 10 December 2018 | Anowarul Amin | No comments yet
In the wake of the human genome project, molecular biology and genetic technologies are tremendously integrating into biomedical research. Currently, PCR, qPCR, and sequencing are key tools in the clinical laboratory for the detection and characterisation of microorganisms and genetic disorders.
Molecular biology also intervenes in changing the human genetic code by recombinant DNA technology that leads to the production of vaccines and drugs.1 Likewise, nucleic acids – for instance, genes, oligonucleotides, and ribozymes – hold promise to be used as therapeutics.2 However, the progress of molecular biology and genetic technology has restricted our understanding of gene organisation, mutation, and expression at the DNA and RNA level. Nevertheless, most diseases are mainly caused by protein malfunctions due to instability, biochemical dysfunction or loss of protein-protein, or protein-DNA/RNA interactions. Therefore, protein interaction and functional networks must be elucidated to comprehensively understand the molecular basis of diseases, which in turn can help identify methods for prevention, diagnosis, and treatment.
Despite research progression, the fundamental role that proteins play in the onset of disease remains obscure. The significant knowledge gaps in the understanding of protein function on disease pathogenesis contribute to the lack of authentic methods for diagnosis and treatment and therefore it has become imperative to develop novel approaches to comprehend the function of the proteins in the context of health and diseases.3 Proteomics is an embracing term mostly used in a fundamental study to describe the large-scale study of the structure and function of proteins. Proteomics offers a powerful set of tools for the direct high-throughput study of gene function. Currently, proteomics is making a key contribution to our understanding of proteins; specifically, how their expression, structure and function can cause diseases. These approaches also allow us to compare the protein profiles of cells in healthy and diseased states that can be used to identify proteins associated with disease development and progression.4
Although limited, novel proteomics techniques such as mass spectroscopy, protein microarray, chromatography, and 2D electrophoresis are accelerating our understanding and knowledge to help us identify the role of proteins in disease regulation and providing us with potential diagnostic and prognostic indicators. Many of the techniques enable the detection of biomarkers for direct medical applications in a range of diseases, including cancer, immune rejection after transplantation, and infectious diseases such as tuberculosis, dengue, and malaria. Recently, proteomics has been used to access branding proteins for therapeutic targets.4-6 The advances of synthetic biology, computational biology, and bioinformatics have significantly accelerated expansion in these areas. As shown in Figure 1, this article briefly outlines current and potential proteomics technologies that broadly solve the mystery of diseases and drugs.
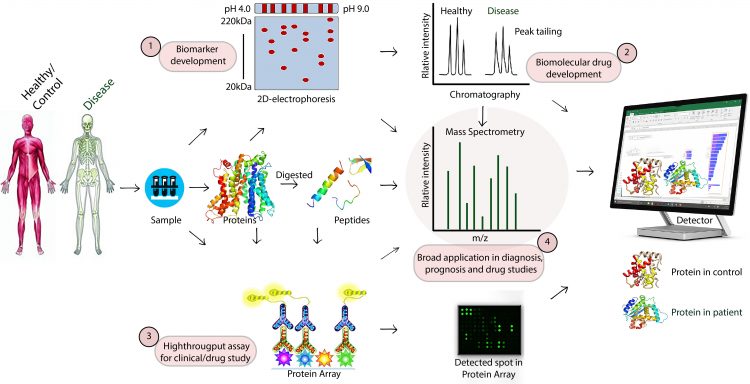

Figure 1: Schematic representation of proteomics techniques. Blood cells or tissues are used for the extraction of total proteins or polypeptides. The desired biomarkers or protein of interests for diseases and therapeutic targets are then detecting by (1) 2-D electrophoresis, (2) chromatography, (3) Protein array, or (4) Mass spectrometry techniques.
1. Two-dimensional (2D) electrophoresis
2D electrophoresis was originally used to analyse mixtures of proteins by two properties in two dimensions, respectively (Figure 1). Although this technology is no longer used in modern proteomics, it still has distinct advantages and key features.7 One of its key features is the unique ability to separate thousands of proteins at once. It provides direct visual confirmation of changes in proteins between healthy vs diseased or treated vs control patients. 2DE gels have been successfully used in biomarker discovery in neurodegenerative diseases such as Parkinson’s disease and cancer biomarkers like gastric carcinoma and breast cancer.8 In addition, 2DE also assists in whole proteome analysis for drug discovery research;8 hence, applications of this technique remain active. However, 2DE is labour-intensive and has a relatively low-throughput, thus it’s mostly used in small-scale basic research studies. The development of the differential imaging gel electrophoresis (DIGE) technique increases the sensitivity and reproducibility of 2DE and hence its application can be perceived in future proteomics.9
2. Chromatography
Pharmaceutical industries have extended their therapeutic products from peptide drugs to large biomolecules such as recombinant proteins or monoclonal antibodies. The production of these important biomolecules frequently requires many purification steps to in order to obtain sufficiently pure products at bulk level. Chromatography is a distinct separation, identification, and purification method often used for determining biomolecular therapeutics in clinical and pharmaceutical industries.10 This method separates small to large-size biomolecules such as proteins and peptides from a complex mixture based on size, shape, charge, and hydrophobic groups. Chromatography techniques characterise antibody-drug conjugates (ADCs) for cancers, atherosclerosis and anti-inflammatory treatments.11 In addition, the pharma and biotechnology industry have used chromatography methods to identify potential vaccines for the prevention of infectious diseases such as dengue, malaria, and chikungunya virus.12 Affinity chromatography, liquid chromatography (LC), high-performance liquid chromatography (HPLC), size exclusion chromatography (SEC), and reversed-phase liquid chromatography (RPLC) are the most frequently used analytical methods for protein characterisation and quality control of ADCs and vaccines in the pharma and biotechnology industries. The development and use of chromatography-based methods for the identification, validation and analysis of biomarkers for diseases have been reviewed. Nevertheless, the first step in the search for biomarkers relies on powerful analytical techniques that enable the separation and identification of thousands of endogenous protein compounds. In this context, chromatographic methods have a limitation, thus chromatography coupled with mass spectrometry appear as methods of choice.
3. Protein microarray
Protein microarray is a term that refers to the miniaturisation of thousands of protein assays on one small plate. It utilises highly specific antigen-antibody recognition to build a protein detection system via the immobilisation of purified proteins on glass slides. Microarray technology has the potential to monitor complex intracellular protein expression and simultaneously study the interaction between proteins and many potential partners in a high-throughput manner. To date, this assay has been developed to detect protein-protein, protein-DNA, protein-RNA, protein-lipid, and protein-drug interactions.13 Thus, protein interaction arrays are also being integrated into drug screening processes by early profiling of drug candidates regarding their efficacy and toxicity. In addition, one of the most relevant applications of protein microarrays is the detection of biomarkers for various diseases, including cancer, where the importance of early detection is fundamental.13 Systematic rheumatic disease and autoimmune diseases can be accurately diagnosed by the presence of nuclear proteins and nucleoprotein complexes as targets for antibodies to detect the phase of diseases. It is widely used for expression profiling in the clinic to understand how pathogens can modify cell pathways; this is a good strategy to establish preventive/therapeutic interventions for infectious diseases.14 However, there are a few limitations that make routine application difficult. For example, different proteins may be coated in differing amounts on the slides, resulting in various levels of fluorescence signal; the assay is therefore not suitable as a quantitative assessment when comparing results between various proteins. It also displays minimal nonspecific binding to minimise background noise in detection systems. The higher the concentration of the detection antibody, the higher the non-specific interaction, which results in an increase in background noise. Therefore, the whole variety of non-specifically captured target proteins is analysed by mass spectrometry.
4. Mass spectrometry (MS)
Mass spectrometry (MS) is extremely powerful and one of the most appreciated sensitive techniques for detection, quantitation, and structure elucidation of several hundred of metabolites to observe the entire cellular picture of proteins.15 During disease condition, protein expression and interaction can change, and structure can be modified in many ways, such as phosphorylation, methylation glycosylation, ubiquitinylation, and acetylation due to size or complexity. The mass spectrometer can capture these altered proteins first by ionising proteins and then subsequently separating them according to their mass to charge ratio (m/z).6 Then, using mass spectrometric reference maps, the customised software platform automatically identifies and quantifies altered proteins within the database (Figure 1). One of the main advantages of MS is the requirement for just a small amount of sample and high-throughput capability in the examination. For instance, only 5–10μL of serum sample is required to detect the protein fingerprint of diseases.16 Also, nowadays, a single droplet of blood can be placed directly on a special surface to bait protein molecules in the blood for unravelling complex protein-protein interactions during disease progression. It has also developed into a useful tool in early pathogenic bacterial and viral proteins identification including intact viruses, mutant viral strains, capsid proteins, and post-translational modifications for early infectious disease control.16
Characterising changes in protein expression between a tumour and healthy tissue, or between the blood of diseased and healthy individuals, is a common approach. Here, mass spectrometry (MS)-based clinical proteomics is providing unbiased biomarker discovery from complex biological samples such as plasma, brain, or liver tissue for tailoring medical decisions of complex diseases like cancer, neurodegeneration, cardiac, and kidney disorder.17 Currently, this technology identifies ~400 biomarkers and allows us to detect patients with lung, breast, thyroid or ovarian cancer with high accuracy.18 The ongoing ambitions to better understand signalling pathways for drug development are being addressed by MS, that will help guide lung, breast, and pancreatic cancer therapy in the future.
Conclusion
The goal of proteomics is to determine the interdependence of cellular processes for abnormal or disease conditions. Current proteomics approaches can provide insights into biological processes at the functional level to determine promising biomarker candidates for diseases and therapeutic targets. In addition, proteomics methods for quantification of proteins in biological samples have been routinely used for the diagnosis of diseases and for monitoring treatment. Although large-scale protein quantification or sensitivity in complex samples remains a challenging task, a large amount of effort has been made to advance the technologies. Additionally, instrument operation, data acquisition and analysis still require highly specialised expertise, but discovering new reagents, novel sampling formats, new analysers and scanning techniques, as well as customised software can help to overcome these limitations.
Biography
Dr Anowarul Amin is currently a Fellow at National Institutes of Neurological Disorders and Stroke (NINDS) of National Institutes of Health (NIH) studying organelles specific diseases. Dr Amin obtained his PhD from the Tokyo University of Science, Japan. Dr Amin’s main research goal is to understand how proteins function as molecular machines in living cells. After his PhD, he joined as a Postdoctoral Research Associate at Texas Tech University Health Sciences Center (TTUSHS) to characterise diseases associated with membrane proteins for potential therapeutics.
References
- Speers DJ. Clinical applications of molecular biology for infectious diseases. Clin Biochem Rev. 2006;27(1):39-51.
- Sridharan K, Gogtay NJ. Therapeutic nucleic acids: current clinical status. Br J Clin Pharmacol. 2016;82(3):659-72.
- Li H, Han J, Pan J, Liu T, Parker CE, Borchers CH. Current trends in quantitative proteomics – an update. J Mass Spectrom. 2017;52(5):319-41.
- Schubert OT, Rost HL, Collins BC, Rosenberger G, Aebersold R. Quantitative proteomics: challenges and opportunities in basic and applied research. Nat Protoc. 2017;12(7):1289-94.
- Hwang H, Hwang BY, Bueno J. Biomarkers in Infectious Diseases. Dis Markers. 2018;2018:8509127.
- Duan G, Walther D. The roles of post-translational modifications in the context of protein interaction networks. PLoS Comput Biol. 2015;11(2):e1004049.
- Sun F, Cavalli V. Neuroproteomics approaches to decipher neuronal regeneration and degeneration. Mol Cell Proteomics. 2010;9(5):963-75.
- Hu S, Loo JA, Wong DT. Human body fluid proteome analysis. Proteomics. 2006;6(23):6326-53.
- Vilasi A, Monti MC, Tosco A, De Marino S, Margarucci L, Riccio R, et al. Differential in gel electrophoresis (DIGE) comparative proteomic analysis of macrophages cell cultures in response to perthamide C treatment. Mar Drugs. 2013;11(4):1288-99.
- Hage DS, Anguizola JA, Bi C, Li R, Matsuda R, Papastavros E, et al. Pharmaceutical and biomedical applications of affinity chromatography: recent trends and developments. J Pharm Biomed Anal. 2012;69:93-105.
- Martin C, Kizlik-Masson C, Pelegrin A, Watier H, Viaud-Massuard MC, Joubert N. Antibody-drug conjugates: Design and development for therapy and imaging in and beyond cancer, LabEx MAbImprove industrial workshop, July 27-28, 2017, Tours, France. MAbs. 2018;10(2):210-21.
- Weaver SC, Osorio JE, Livengood JA, Chen R, Stinchcomb DT. Chikungunya virus and prospects for a vaccine. Expert Rev Vaccines. 2012;11(9):1087-101.
- Sutandy FX, Qian J, Chen CS, Zhu H. Overview of protein microarrays. Curr Protoc Protein Sci. 2013;Chapter 27:Unit 27 1.
- Jean Beltran PM, Federspiel JD, Sheng X, Cristea IM. Proteomics and integrative omic approaches for understanding host-pathogen interactions and infectious diseases. Mol Syst Biol. 2017;13(3):922.
- Altuntas E, Schubert US. “Polymeromics”: Mass spectrometry based strategies in polymer science toward complete sequencing approaches: a review. Anal Chim Acta. 2014;808:56-69.
- Chandramouli K, Qian PY. Proteomics: challenges, techniques and possibilities to overcome biological sample complexity. Hum Genomics Proteomics. 2009;2009.
- Crutchfield CA, Thomas SN, Sokoll LJ, Chan DW. Advances in mass spectrometry-based clinical biomarker discovery. Clin Proteomics. 2016;13:1.
- Liang SL, Chan DW. Enzymes and related proteins as cancer biomarkers: a proteomic approach. Clin Chim Acta. 2007;381(1):93-7.
Related topics
Analytical Techniques, Protein, Proteomics