How do fast-growing bacteria resist antibiotics?
Posted: 13 June 2022 | Ria Kakkad (Drug Target Review) | No comments yet
A new study has revealed how some fast-growing bacteria could be retreated with antibiotics.


In a new study published in eLife, scientists have demonstrated how some fast-growing bacteria can be retreated with antibiotics. The results show that fast-growing individuals within bacterial colonies display a significantly higher expression of active ribosomes – particles within the cell that synthesise proteins. This helps the bacteria avoid the accumulation of an important class of antibiotics called macrolides and therefore resist therapies. These findings could be used to inform the development of improved antibiotic compounds that target this survival strategy.
For an antibiotic to be effective against infection, it needs to reach its cellular target at a sufficient concentration to inhibit bacterial growth.
“Antibiotic resistance continues to threaten the viability of current treatments. We need to understand how individual bacteria within a colony can prevent antibiotics from entering their cells, so that we can target this mechanism with new therapies,” said Urszula Łapińska, PDRA at the University of Exeter, UK. “Most existing data around drug permeability in bacteria has been attained through measurements that either take an average result from a large population or are derived from a small number of bacteria. This means that little is known about the variability in individual drug accumulation across many single cells in a bacterial colony.”
To address this gap, the team began with the hypothesis that variations in how bacteria respond to drugs could be driven by the varying drug transport rates between individual cells. To test this, the team used a multi-analytical approach, combining microfluidics-microscopy, bacteria that pose a health threat – namely E. coli, Pseudomonas aeruginosa, Burkholderia cenocepacia and Staphylococcus aureus and fluorescent probes derived from antibiotics by researchers at the University of Queensland, Australia. This approach allowed the team to examine the interactions between commonplace antibiotics and many live, individual bacteria in real-time, during drug dosage. By combining this approach with mathematical modelling techniques, the team obtained data that they could use to quickly and efficiently identify individual bacteria that are antibiotic resistant.
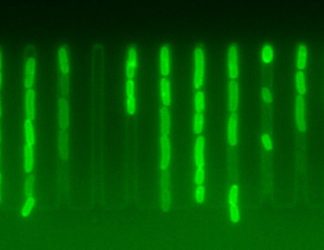

Individual cells display dramatically different accumulation of antibiotics
[Credit: Ula Łapińska (CC BY 4.0)]
Their analyses demonstrated that fast-growing individuals within a colony avoid the accumulation of macrolides within their cells – a discovery that contrasts with current thinking that slow cell growth is the major contributor to survival to antibiotics without genetic variation. This avoidance is enabled by a significantly higher number of ribosomes before drug treatment, compared to the individuals’ slow-growing counterparts. Ribosomes enable essential cellular processes, including efflux – a system that pumps toxic substances, such as antimicrobial compounds, out of the cell.
Using this new knowledge, the researchers then showed that chemically manipulating the outer membrane of the bacterial cells allows to eradicate fast-growing variants that exhibit low macrolide accumulation, therefore aiding our fight against antibiotic resistance.
“This work reveals a hitherto unrecognised survival strategy in some members of bacterial colonies,” concluded Dr Stefano Pagliara, Senior Lecturer in Microfluidics at the University of Exeter. “These insights will directly benefit microbiologists and clinicians who are working on the development of more effective antibiotic therapies. In the longer term, we hope that using our novel approach in clinical settings will help inform the design of improved drugs and help us in the fight against antibiotic resistance.”
Related topics
Antibiotics, Microscopy
Related organisations
Queensland University, University of Exeter
Related people
Dr Stefano Pagliara, Urszula Łapińska