Harnessing the power of genomic sequencing augments diagnosis and treatment of lymphoid cancer
Posted: 9 February 2018 | Drug Target Review | No comments yet
A new study has established that hybrid-capture sequencing is the method of choice for sequencing “actionable” gene mutations across the most common forms of lymphoid cancer.
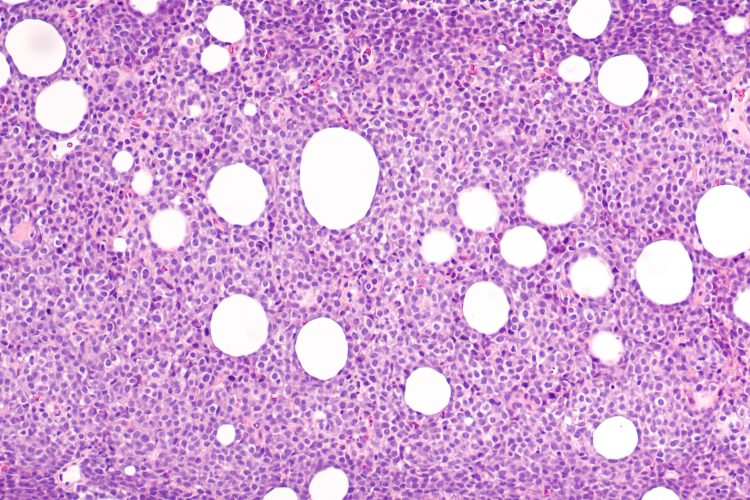
Due to its applicability in routinely acquired formalin-fixed, paraffin-embedded tissues, this assay can be implemented by clinical laboratories into routine diagnostic workflows. It reliably identifies potentially actionable gene mutations in 91 per cent of patients. This assay will bring the benefits of precision diagnosis and individualised therapy to patients with lymphoid cancer.
“To realise the benefits of the most recent progress in cancer genomics, clinical implementation of precision medicine approaches is needed in the form of novel biomarker assays. Fully implemented targeted sequencing-based assays in routine diagnostic pathology laboratories are currently lacking in lymphoid cancer care,” explained Christian Steidl, MD, Senior Scientist at the BC Cancer and Associate Professor in the Department of Pathology and Laboratory Medicine, University of British Columbia, Vancouver, Canada. “Our findings demonstrate the feasibility and outline the clinical utility of integrating a lymphoma-specific pipeline into personalised cancer care.”
Lymphoid cancers include diffuse large B-cell lymphomas (DLBCL), follicular lymphoma (FL), and chronic lymphocytic leukemia (CLL), which were the focus of the present study. In recent years, several sequencing-based assays have been developed, but their clinical applicability and utility for patients with specific mutations still needs to be shown. Actionable gene mutations are defined by having a reported correlation with treatment outcome, a molecular association with a particular targeted therapy, or increasing pathologic-diagnostic accuracy. Currently, most genetic mutations associated with lymphoid cancers have been identified.
“This study is remarkably comprehensive, which will help any molecular laboratory design and implement their own next-generation sequencing lymphoma panel using this work as a template,” commented Robert S. Ohgami, MD, PhD, Assistant Professor of Pathology at the Stanford University Medical Center, in an accompanying editorial.
Unlike whole-genome sequencing, this assay targets a select set of genes or regions with known associations with lymphoid cancer, allowing for more rapid detection of a variety of mutations. According to this study, capture hybridization is a better method over amplicon-based sequencing, when using formalin-fixed, paraffin-embedded tissue.
Also, this assay is continuously modifiable due to its modular flexible design. The 32-gene next-generation sequencing panel described in the paper was developed with input from a group of six specialists who kept updating it based on the latest available information. “This allows for continuous integration of additional gene features as our knowledge base improves,” added Dr. Steidl.
The investigators applied the assay to tissues from 219 patients with lymphoid cancers (114 FL, 76 DLBCL, and 29 CLL) who were treated in British Columbia, Canada, between 2013 and 2016. At least one actionable mutation was found in 91 per cent of the tumours. The assay revealed cancer subtype–specific mutational profiles that were highly similar to published mutational profiles for all three types of lymphoid cancer. The assay was also found to have 93 per cent concordance with whole-genome sequencing.
“Our developed assay harnesses the power of modern sequencing for clinical diagnostics purposes and potentially better deployment of novel treatments in lymphoid cancers. We believe our study will help establish evidence-based approaches to decision making in lymphoid cancer care. The next steps are to implement sequencing-based biomarker assays, such as reported in our study, in accredited pathology laboratories. Toward the goal of biomarker-driven clinical decision making, testing of potentially predictive biomarker assays is needed alongside clinical trials investigating novel cancer therapeutics,” noted Dr. Steidl.
The study is published in The Journal of Molecular Diagnostics.
Related topics
Assays, Biomarkers, Genetic Analysis, Genomics
Related conditions
lymphoid cancer
Related organisations
Stanford University Medical Center, The University of British Columbia
Related people
Dr Christian Steidl, Dr Robert S. Ohgami