New software could enable the design of DNA robots for drug delivery
Posted: 20 April 2021 | Victoria Rees (Drug Target Review) | No comments yet
Researchers have developed software that can design complex DNA nanodevices which could be used to deliver medicine while in the body.


Researchers at Ohio State University, US, have developed a new tool that can design much more complex DNA robots and nanodevices than were possible before in a fraction of the time. According to the scientists, tiny DNA-based robots and other nanodevices could be used to deliver medicine inside our bodies, detect the presence of deadly pathogens and help manufacture increasingly smaller electronics.
The team unveiled the new software called MagicDNA. They say that this will help researchers design ways to take tiny strands of DNA and combine them into complex structures with parts like rotors and hinges that can move and complete a variety of tasks, including drug delivery.
“Now, nanodevices that may have taken us several days to design before now take us just a few minutes,” said Associate Professor Carlos Castro, co-author of the study.
“Previously, we could build devices with up to about six individual components and connect them with joints and hinges and try to make them execute complex motions,” said study co-author Professor Hai-Jun Su. “With this software, it is not hard to make robots or other devices with upwards of 20 components that are much easier to control. It is a huge step in our ability to design nanodevices that can perform the complex actions that we want them to do.”
The team say the software has a variety of advantages that will help scientists design better, more helpful nanodevices and hopefully shorten the time before they are in everyday use.
One advantage is that it allows researchers to carry out the entire design truly in three-dimensions (3D). Earlier design tools only allowed creation in two-dimensions (2D), forcing researchers to map their creations into 3D. That meant designers could not make their devices too complex.
The software also allows designers to build DNA structures “bottom up” or “top down.” In bottom up design, researchers take individual strands of DNA and decide how to organise them into the structure they want, which allows fine control over local device structure and properties. However, they can also take a top down approach where they decide how their overall device needs to be shaped geometrically and then automate how the DNA strands are put together.
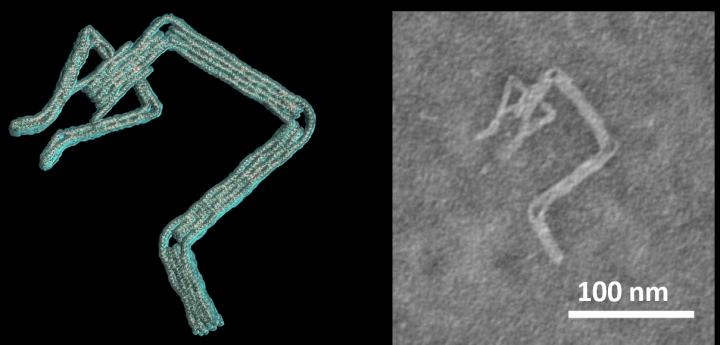

A robot arm nanodevice with claw that can pick up smaller items [credit: Ohio State University].
Combining the two allows for increasing complexity of the overall geometry while maintaining precise control over individual component properties, the team say.
Another key element of the software is that it allows simulations of how designed DNA devices would move and operate in the real world.
“As you make these structures more complex, it is difficult to predict exactly what they are going to look like and how they are going to behave,” Castro said. “It is critical to be able to simulate how our devices will actually operate. Otherwise, we waste a lot of time.”
As a demonstration of the software’s ability, co-author Anjelica Kucinic, led the researchers in making and characterising many nanostructures designed by the software.
Some of the devices they created included robot arms with claws that can pick up smaller items. The researchers suggest that, in the future, they may be able to design a DNA robot that, after injection into the bloodstream, can detect a certain pathogen.
“But a more complex device may not only detect that something bad is happening, but can also react by releasing a drug or capturing the pathogen,” Castro said. “We want to be able to design robots that respond in a particular way to a stimulus or move in a certain way.”
The study was published in Nature Materials.
Related topics
DNA, Drug Delivery, Informatics, Lab Automation, Nanomedicine, Robotics
Related organisations
Ohio State University
Related people
Anjelica Kucinic, Associate Professor Carlos Castro, Professor Hai-Jun Su