Genes behind antibiotic activity in bacteria uncovered by metabologenomics
Posted: 2 August 2021 | Anna Begley (Drug Target Review) | No comments yet
Researchers have used comparative metabologenomics to uncover what may be “silencing” bacteria to produce desirable compounds.
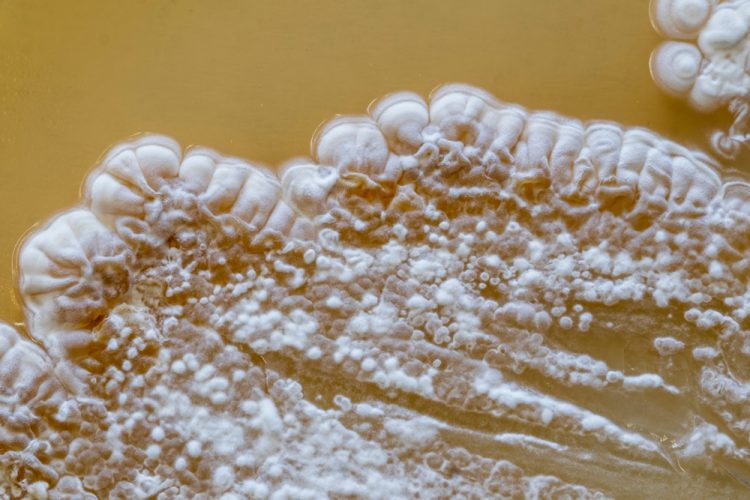


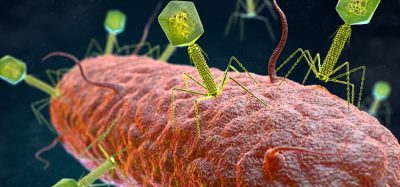
A team of scientists at the Washington University in St Louis, US, have utilised the benefits of comparative metabologenomics to try to uncover what may be “silencing” Streptomyces bacteria and preventing it from producing desirable compounds encoded by its genes. Drug-like molecules are hidden in the genomes of Streptomyces bacteria, however isolating those genomes has proved challenging. According to the team, metabologenomics may therefore be the key to identifying potential antibiotic producers.
“We examined genetic differences across the genomes of Streptomyces while at the same time looking at antibiotic outputs,” said corresponding author Joshua Blodgett. “This study highlights comparative metabologenomics as a powerful approach to expose the features that differentiate strong antibiotic producers from weaker ones.”
Blodgett’s team compared a group of antibiotic-producing strains of Streptomyces and other non-producing or poor-producing strains to reveal genomic differences that could affect drug production. They found a few key differences between the strains. Notably, good producers of polycyclic tetramate macrolactam (PTM) antibiotics seemed to benefit from griseorhodin production which the researchers did not anticipate and initially had tried to eliminate.
Comparative metabologenomics also revealed that the presence or absence of two to three nucleotides that can tune the switches that drive PTM into antibiotic production. Previously, this type of fine control had been found in certain bacteria that cause disease but had largely been overlooked in bacteria that produce drugs.
“Our work highlights the problem of silent gene clusters and the need to understand them for next-generation drug discovery,” concluded Blodgett. “Comparative metabologenomics is a generally adoptable strategy and we hope that others might use to examine their own strains and drug pathways.”
The study was published in PNAS.
Related topics
Analytical Techniques, Antibiotics, Genomics, Metabolomics, Microbiology
Related organisations
Washington University in St. Louis
Related people
Joshua Blodgett