Farther than fat: how lipids change research
Posted: 8 May 2019 | Dr Christian Klose -Head of Research & Development at Lipotype GmbH | No comments yet
Today we are beginning to realise that lipids have been inadequately represented in biological and medical research. For decades, studies in the fields of DNA and proteins have been pushed but despite these adverse conditions, lipid research has made pioneering discoveries over the course of the last seven years. These findings have facilitated better appreciation of the importance of fat molecules and the reason for that is a paradigm shift in lipid research.
LIPIDS are a vastly influential class of molecules. Alzheimer’s, diabetes and cancer are all known to be influenced by lipids; but they are a vital part of our diet and skin structure too. Thus, industry and academic research are dependent on lipid analysis to leverage their explanatory power. However, the lipid research methods of the past have hindered swift progress for decades, with inaccuracy, lengthy analysis time and high prices suppressing promising research projects from gaining knowledge and developing applications. However, change came with technological breakthrough.
Prior to the new millennium, lipid analysis was mainly achieved through chromatography methods, which was both time- and cost-intensive. Only lipid classes could be analysed and consequently the whole spectrum of thousands of lipid species was ignored. Since then, technological development changed its direction to focus on mass spectrometry (MS)-based methods and the subsequent breakthroughs of the last two decades led to shotgun lipidomics. Shotgun lipidomics technology solves the problems of the past and now provides insights relevant to medical and biological research. Today, lipids and lipid compositions can be analysed precisely and swiftly1 and researchers receive answers to questions they had not even asked before. Lipid research has become lipidomics.
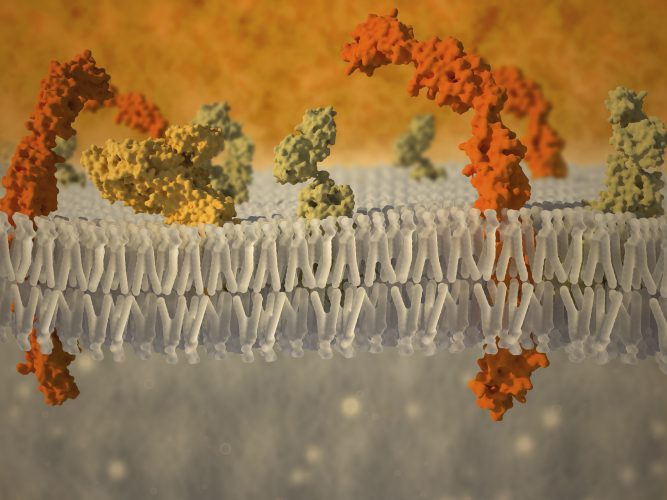
Cell membranes are the key element of life. They separate the cell’s inside from its outside to create an enclosed, protected environment and consist of a lipid bilayer with embedded proteins.
In analogy to genomics, lipidomics is the study of lipids and lipid metabolites in biological systems. Targeting thousands of chemically-distinct lipid molecules, lipidomics unveils the many different biological functions of lipids, which range from storing energy, serving as hormones and signalling molecules for the dynamic matrix of the cell membrane. Typically, lipidomics is applied to analyse the lipid composition of sets of biological samples of different types in basic research, or to compare two or more cohorts from healthy subjects and from patients, treated and untreated, healthy and diseased, female and male.
In contrast to most metabolomics approaches,lipidomics can now cover the whole spectrumof lipids with one class-specific standard perlipid class. Only signalling lipids require otherapproaches. Thus, the shotgun platform deliversquantitative lipidomic analysis, surpassingcomparative experimental setups that provideonly relative quantitation.
Lipidomics used to be considered a subgenre ofmetabolomics. However, due to its rapid increase in popularity and applicability, lipidomics acquired independence and today is often quoted as the fastest growing omics science. The joint efforts ofMS method development, MS data bioinformatics, lipid-specific biostatistics and hands-on biological and medical research have paid off.
Perhaps the biggest breakthrough in lipidomicscame when academia and industry started tofollow what lipid researchers had advocated fordecades: fat is not just fat, it is thousands of activemolecules. Lipids started to emerge as playersin a broad spectrum of biological processes.
Companies and research groups, which had previously had no association with lipids, began to encounter these molecules and were faced with questions like “How do I analyse what lipids my samples are comprised of?” or “Does my drug challenge the patient’s state of lipid metabolism?”.
Recent research results conclude that specificlipid signatures obtained by shotgun lipidomicscould help to diagnose diseases in some cases evenyears prior to the onset of traditional symptoms.
In Scandinavia – specifically Malmö and Lund in Sweden, and Helsinki in Finland – this mindset has already led to pioneering research.
The lipidomic analysis of the 2,000 patients in the Malmö Diet and Cancer Study tackled the unknown molecular details of the generally known association of cardiovascular diseases and diabetes with dyslipidemia. They discovered a lipid signature linking reduced levels of specific unsaturated fatty acids in complex lipids with diabetes and myocardial infarction. This emphasises the importance of lipids in the molecular pathogenesis of both diseases.2
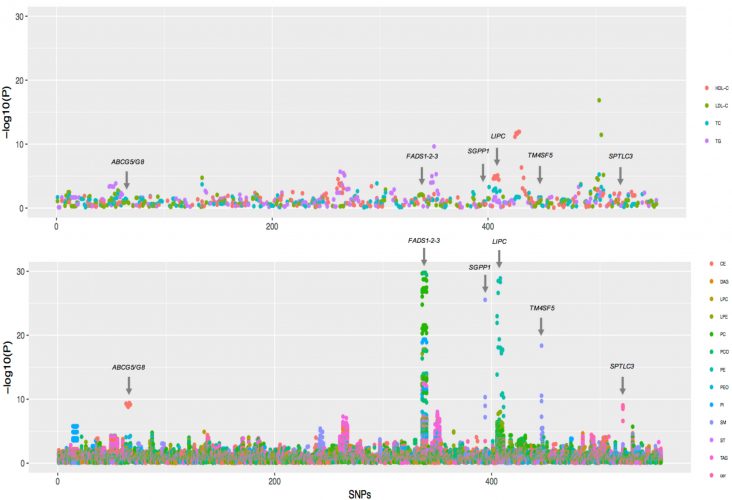
Explanatory power of traditional lipid measurements and lipidomics analysis in comparison. Association of known single nucleotide polymorphisms for lipids with traditional lipid measurements (upper panel) and lipidomics analysis data (lower panel). The data clearly shows that molecular lipidomic data have much stronger association (lower P values). Noticeable associations have been annotated.3
In Helsinki, a multi-omics approach of genomics and lipidomics combined genomic risk prediction scores for a variety of diseases with lipidomics phenotyping. They discovered that despite dietary influences on the blood plasma, lipid composition, specifically polyunsaturated lipid species levels, feature a strong genetic sharing. Furthermore, they compared the capacity to discover lipid-associated genetic variances of lipidomic data and traditional clinical lipid measurements, the current standard in clinical diagnostics for cardiovascular disease prevention. The approach clarified that molecular lipidomic data trumps traditional lipid measurements by several orders of magnitude and provides the tools to capture individual or collectives of lipids as potent independent disease risk factors.3
Both studies highlight blood sample lipids as a potent resource to identify cardiovascular diseases early on. However, many metabolic disorders, as well as several neurodegenerative diseases, have already been linked to lipid metabolism, with new diseases continuously being added. As such, preventive medicine is harnessing lipidomics as a tool of great explanatory power, which furthermore increases continuously as the number of available lipid signatures grows. In the next decade, shotgun lipidomics could become a daily clinical routine and transform how the general public experiences medicine.
However, the applicability of lipidomics is not limited to identifying lipid signatures in clinical diagnostics and biotech. Mode of action studies, support of intervention studies, drug discovery and development, pharmacokinetic or medicinal efficiency evaluation, as well as many other areas of pharma research, are joining the lipid revolution.
The lipid OHOA is an example of that. This fatty acid is used in membrane lipid therapy for the treatment of glioma and other solid tumours. The anti-tumour effect was thought to be mediated by affecting biophysical membrane properties. Lipidomic analysis enlightened the influence of OHOA on the molecular level. Adding this lipid to the drug led to its incorporation into several lipid classes and stimulated a complex response of multiple signalling pathways and intracellular proteins to facilitate the transport of the active ingredient. Thus, the mode of action study revealed that OHOA, as a known membrane-properties changer, not only affects cancer signalling transduction cascades but also intracellular transport.4
Lipidomics is not just of interest in basic science or clinical diagnostics. The development of nutraceuticals and functional foods, as well as testing cosmetics and dermatological products, are fields of applications. The spectrum of current research and industrial product development reflects the impact of lipidomics on all areas of life sciences, further consolidating lipidomics as a tool of equivalent power next to genomics and proteomics in industry and academia. Nonetheless, this is only the beginning. The lipidomics field has much work to do.
Today’s lipidomics is the outcome of innovation in technology and methodology development. It is a global project that stems from the pioneers and hubs of lipid research spread all over the world. Though this represents a solid basis for building an inclusive global lipidomics community, the weakness is that there is no standardisation of protocols and software. Almost every lab employs a slightly different method and a slightly different protocol. The consequences are highlighted in an interlaboratory study of 2017, which is a first attempt to sort the good, from the bad and the ugly.5
The next big leap will be to focus on standardised protocols across labs. This will foster progress in the life sciences as lipidomic data will become comparable across different labs, which in turn increases the overall quality of the data. A selection of thought leaders has already taken on the challenge and founded the Lipidomics Standards Initiative (LSI). We are only now beginning to grasp the importance of lipids in biological and medical research. The comprehensive indexing of the biological functions of fat molecules is a Herculean task, but lipids will be worth the effort and lipidomics provides the key to understanding them. We hope that this pursuit will deliver tools for a better life.
Biography
DR CHRISTIAN KLOSE: After receiving his PhD on membrane biology and biophysics, he did a post-doc on mass spectrometry-based lipidomics with Andrej Shevchenko. Christian has published numerous papers in the field of membrane biology and lipidomics. As Head of R&D at , Christian enjoys developing novel technology and putting it into action.
References
- Surma M, et al. “An automated shotgun lipidomics platform for high throughput, comprehensive, and quantitative analysis of blood plasma intact lipids.” European Journal of Lipid Science and Technology 2015, doi.org/10.1002/ejlt.201500145
- Kjellqvist S, et al. “Identification of Shared and Unique Serum Lipid Profiles in Diabetes Mellitus and Myocardial Infarction.” Journal of the American Heart Association 2016, doi.org/10.1161/ JAHA.116.004503
- Tabassum R, et al. “Linking genome to phenome through lipidomic profiles identifies disease susceptibility genes.” American Society of Human Genetics 68th Annual Meeting 2018
- Torgersen ML, et al. “The anti-tumor drug 2-hydroxyoleic acid (Minerval) stimulates signaling and retrograde transport.” Oncotarget 2016, doi. org/10.18632/oncotarget.13508
- Bowden JA, et al. “Harmonizing lipidomics: NIST interlaboratory comparison exercise for lipidomics using SRM 1950-Metabolites in Frozen Human Plasma.” Journal of Lipid Research 2017, doi.org/10.1194/ jlr.M079012
Related topics
Analytical Techniques, Lipidomics
Related organisations
Merck, MilliporeSigma
Related people
Andrej Shevchenko