Image-based characterisation of cancer biotherapeutics
Posted: 17 June 2019 | Laura Murch, Siobhán Leonard, Victoria Juskaite | No comments yet
Scientists from LifeArc, an independent medical research charity in the UK, explain how they’re using the latest imaging technologies to progress drug discovery campaigns and identify the most promising candidates for antibody-based therapies.
Therapeutic antibodies have progressively changed the paradigm of cancer care in recent years, with at least 28 monoclonal antibodies1 and four antibody-drug conjugates (ADCs)2 in clinical use to date. The exponential growth of this field has coincided with significant advancements in imaging platforms, facilitating in-depth pre-clinical assessment of novel therapies. LifeArc has experience in the successful humanisation of therapeutic antibodies (Keytruda, Entyvio, Actemra and Tysabri) and has harnessed the capabilities of kinetic and high-content image analysis systems to identify lead candidates and characterise their impact on physiologically relevant two-dimensional (2D) and three-dimensional (3D) in vitro models. Kinetic platforms offer a powerful method to dissect basic biology, which is crucial to the advancement of promising anti-cancer agents as it enables dynamic changes in cell viability, migration, invasion, immune cell proliferation and immune cell killing to be rapidly assessed in 96- and 384-well formats. High-content systems provide a means to further investigate complex scientific questions with sophisticated sensitivity and, when coupled with intuitive analysis workflows, this intricate, multiparametric data can have a significant impact on the progression of drug discovery programmes.
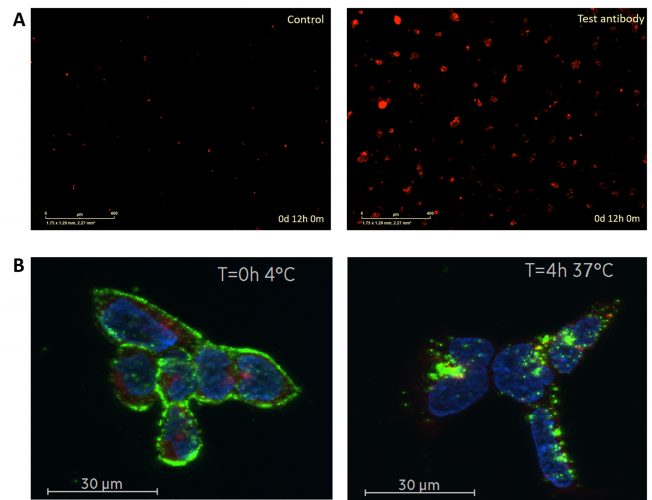
Figure 1: Antibody internalisation in HEK293 cells overexpressing a receptor of interest. (A) An isotype control or a test antibody against the target of interest was incubated with IncuCyte Human FabFluor-pH Red Antibody Labeling Reagent before addition to cells, then monitored over time for red object area on the IncuCyte S3 Live-Cell analysis system using a 10x objective. The labelling reagent only emits a red fluorescent signal at low pH and thus indicates antibody internalisation to the acidic lysosome. (B) Cells stained with an AlexaFluor488-conjugated antibody against the target shown in green, lysosomes stained with Lysotracker Deep Red shown in red and nuclei stained with Hoechst 33342 shown in blue. The antibody appears restricted to the cell membrane at the initial time point (0 hours at 4°C); after a 4-hour incubation at 37°C clear internalisation is observed with the appearance of punctate structures which co-localise with lysosomes.
Antibody internalisation
The majority of successful therapeutic antibodies for oncology are selected for their capacity to induce antigen-dependent cytotoxicity or to provoke immune-mediated tumour cell death. Critical to the success of such mechanisms is antigen availability. Investigating antibody internalisation achieves deeper understanding of antibody and antigen turnover, which allows the best possible drug candidates to be progressed. In addition, ADCs, which enable highly specific delivery of a toxic payload to a target tumour cell, are emerging as effective therapeutics in oncology. The development of robust methodologies that screen for candidate internalisation to the lysosome, where the linker will be cleaved to release the attached cytotoxin, are also essential to an ADC programme. While flow cytometry can be used to monitor antibody internalisation, this method proves difficult to miniaturise and optimise for screening. In order to facilitate high-throughput analysis of hybridoma supernatant and humanised variant internalisation, we routinely utilise an antibody labelling reagent, which emits a fluorogenic signal when processed via acidic lysosomes and endosomes. The antibody/labelled complex can be directly added to cells and monitored in real time to provide vital kinetic data in a rapid manner (Figure 1A). While providing robust results, this information also allows more complex analysis of the intracellular trafficking of candidate antibodies using high-content imaging platforms to be conducted with an informed approach (Figure 1B). We collaborate closely with academics to develop project-specific assays for high-throughput analysis in order to progress drug discovery campaigns. The coupling of kinetic and high-content imaging systems in the screening of antibodies expedites the process of candidate selection by presenting an in-depth picture of intracellular biology and provides further confidence in potential therapeutics. Given the high costs of attrition in the clinic, it has become increasingly important to expand our knowledge of modulators for cancer therapy, and while an understanding of antibody engagement with the target antigen is critical to development, the impact of the Fc domain of a potential biotherapeutic on immune effector functions should also be evaluated.
Fc effector function
Interrogating the potential impact of immune cell effector function includes screening candidates for activation of monocyte or macrophage engulfment of target cells (antibody-dependent cellular phagocytosis (ADCP)), NK cell-mediated target cell death (antibody-dependent cellular cytotoxicity (ADCC)) and activation of the complement cascade to induce target cell death (complement-dependent cytotoxicity (CDC)). In order to set up physiologically relevant assays for candidate screening, Fc effector function assays typically require the isolation and use of primary cells or sera from human blood. The isolation and culture of such material, however, is time consuming and the relatively small cell numbers procured from such isolations can present limitations when setting traditional endpoint assays such as flow cytometry analysis. Assay readouts, such as luminescent cell viability dyes, have further drawbacks; for instance, being unable to distinguish between target cell and effector cell death. In contrast, kinetic imaging systems enable real-time monitoring of co-culture assays and can be multiplexed with more traditional approaches to maximise the insight that can be garnered from a single experiment. Additionally, these assays are typically amenable to miniaturisation, which further increases the output from a single primary cell isolation. In the course of our work, we have focused on developing screening assays for ADCP, ADCC and CDC Fc effector function. The use of apoptosis assay reagent allows for label-free analysis of cell death to be monitored in real time. We routinely combine this with tumour cell-line labelling to allow specific tumour cell death to be analysed. These dyes negate the use of cell perturbing or labour-intensive cell labelling steps for target cells ahead of assay set up. For analysis of ADCP by monocyte-derived macrophages, we have developed a 384-well plate assay to visualise target cells being engulfed by macrophages (Figure 2). This allows specific tumour cell death to be analysed and can be multiplexed with other endpoint viability analyses. Given the feasibility of high-throughput screening versions of ADCC, ADCP and CDC assays using kinetic imaging platforms, it is now possible to gain insight into the Fc effector function at earlier stages of development for larger numbers of candidates using physiologically relevant cell types. As such, this increases our ability to identify lead candidates with profiles that suit the intended mode of action in the clinic. Impact on the immune cell compartment is not limited to Fc effector function, but also includes interrogating the effect of potential therapies on T-cell proliferation as well as tumour cell-killing assays.
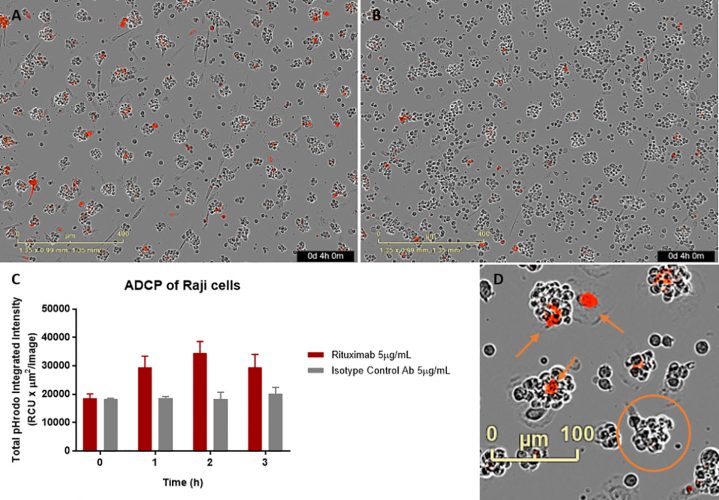
Figure 2: Raji target cells labelled with IncuCyte pHrodo Red Cell Labelling Kit were pre-treated with 5μg/mL Rituximab (A) or control antibody (B) and subsequently seeded onto a layer of primary human monocyte-derived macrophages. Plates were incubated on the IncuCyte S3 Live-Cell analysis system and images taken each hour using the 10x objective. Example images shown after 4-hour incubation for Rituximab treated (A) and isotype control treated (B) conditions. ADCP activity was quantitated by measuring the total pHrodo red fluorescence intensity in macrophage cells and plotted for comparison over time (C). It was noted that the target Raji cells form small clusters over time, as they would during routine culture (highlighted in orange circle (D)). The monocyte‑derived macrophages interact with, migrate over and phagocytose the target cells (examples of this highlighted by orange arrows in (D)).
T-cell activity
In addition to assessing antibody internalisation and Fc effector functions, LifeArc routinely utilises imaging for kinetic image-based analysis of effector T-cell function. Effector T-cells have the potential to control tumours, and therapies that boost T-cell killing of tumour cells have been/are currently in development, including immune checkpoint inhibitors, adoptive T-cell therapy and chimeric antigen receptors. Development of such therapies requires establishment of robust assays to measure effector T-cell activity, which can be analysed by assessment of cell proliferation, cytokine release, expression of cell-surface markers and tumour cell-killing ability. While these functions may be analysed by conventional flow cytometry methods, imaging offers unique advantages, such as visualisation of T-cell morphological changes including clustering as well as the possibility of multiplexing with endpoint assays. Imaging-based methods are particularly useful to assess therapeutic effects on the immune cell-killing of tumour cells, which is often analysed by co-culturing tumour cells with peripheral blood mononuclear cells (PBMCs) or cytotoxic T-cells. This requires tumour cells to be distinguishable from immune cells, which can be achieved by using fluorescently labelled tumour cells in imaging assays. We have established kinetic image-based immune cell-killing assays by using stable tumour-cell lines expressing nuclear-restricted green fluorescent protein (GFP). Co-culture of these cells with activated cytotoxic T-cells clearly demonstrates reduced tumour cell growth (Figure 3A). In addition to this, the assay may be combined with cytotoxicity or apoptotic reagents for multiplexed measurement and automatic quantitation of tumour-cell apoptosis or cytotoxicity in a single well. While 2D cell cultures are valuable, this system does not recapitulate the physiological 3D environment and as a result sometimes provides ambiguous and non-predictive data for in vivo responses, which is a contributory factor to drug failure in clinical trials.3 We measured T-cell mediated destruction of tumour spheroids with our assay (Figure 3B), which, as with monolayer co-culture assays, may be combined with apoptosis or cytotoxicity reagents.
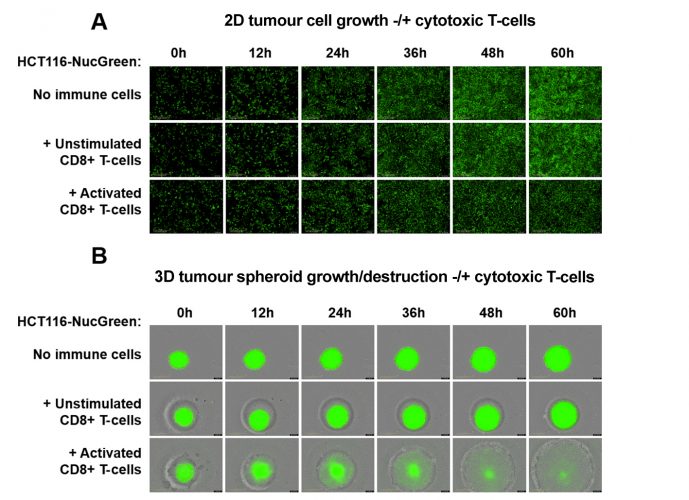
Figure 3: Monitoring tumour cell or spheroid growth in immune/tumour cell co-culture assays. (A) Tumour cell growth is reduced by activated cytotoxic T-cells in a co-culture assay. HCT116 cells stably expressing the nuclear-restricted green fluorescent protein (GFP) (HCT116-NucGreen) were plated alone or in the presence of unstimulated or activated cytotoxic CD8+ T-cells. Tumour cell growth (shown in green) was monitored for 60 hours. (B) Tumour spheroids are destroyed by activated cytotoxic T-cells. HCT116-NucGreen spheroids were incubated alone or in the presence of unstimulated or activated cytotoxic CD8+ T-cells. Tumour spheroid growth or destruction (shown in green) was monitored for 60 hours. Images taken in brightfield and green channel. Note the presence and proliferation of activated immune cells in the periphery of the spheroid. Images were taken using a 10x objective.
Summary
The use of kinetic and high-content imaging platforms to characterise biotherapeutics provides enhanced insight into therapy effects, particularly when paired with physiologically relevant models and 3D systems. Imaging assays deliver a wealth of content through multiparametric analysis of cell number, morphology and area, in addition to fluorescent readouts. Furthermore, these insightful assays can often be multiplexed with more ‘standard’ assay formats; for example, flow cytometry for effector/target-cell analysis or testing supernatants for cytokine release profiles in ADCC and T-cell killing. The ability to miniaturise many of these delicate live cell assays without sacrificing data quality provides a powerful means to accelerate drug discovery campaigns. Coupled with the advancements in automation, imaging-based approaches in combination with established endpoint assays enables creation of an intricate cellular picture of each candidate at a much earlier timepoint from similar numbers of experiments. Ultimately, the routine use of imaging technologies in primary screening campaigns will lead to better characterised biotherapeutics being selected as lead candidates for cancer therapy.
Biographies
 SIOBHÁN LEONARD joined LifeArc in 2016 as a Scientist within the Target Validation Group in the Centre for Therapeutics Discovery. Siobhán gained a BSc in Biomedical Science, a PhD in Cellular and Molecular Biology and carried out a post-doctoral fellowship at the National University of Ireland, Maynooth. With eight years experience in early-stage drug discovery, Siobhán is involved in designing cell-based assays to support the pre-clinical development of antibody, antibody drug-conjugate and small molecule programmes at LifeArc.
SIOBHÁN LEONARD joined LifeArc in 2016 as a Scientist within the Target Validation Group in the Centre for Therapeutics Discovery. Siobhán gained a BSc in Biomedical Science, a PhD in Cellular and Molecular Biology and carried out a post-doctoral fellowship at the National University of Ireland, Maynooth. With eight years experience in early-stage drug discovery, Siobhán is involved in designing cell-based assays to support the pre-clinical development of antibody, antibody drug-conjugate and small molecule programmes at LifeArc.
 LAURA MURCH is a Scientist within the Target Validation Group at LifeArc and is responsible for designing cell-based assays to characterise and validate biotherapeutic and small molecule candidates for various drug discovery programmes. Laura gained a BSc in Human Biology, a PhD in Leukaemia Biology, and carried out a post-doctoral fellowship in Rheumatoid Arthritis at the University of Birmingham. Laura has worked in R&D for nine years and prior to joining LifeArc in 2018 held posts at MedImmune Ltd and Crescendo Biologics Ltd.
LAURA MURCH is a Scientist within the Target Validation Group at LifeArc and is responsible for designing cell-based assays to characterise and validate biotherapeutic and small molecule candidates for various drug discovery programmes. Laura gained a BSc in Human Biology, a PhD in Leukaemia Biology, and carried out a post-doctoral fellowship in Rheumatoid Arthritis at the University of Birmingham. Laura has worked in R&D for nine years and prior to joining LifeArc in 2018 held posts at MedImmune Ltd and Crescendo Biologics Ltd.
 VICTORIA JUSKAITE is a Scientist within the Target Validation Group at LifeArc, located at the Stevenage Bioscience Catalyst science park. Victoria obtained her BSc in Biomedical Science, followed by an MRes in Biomedical Research and PhD in Molecular Cell Biology at Imperial College London. After completion
VICTORIA JUSKAITE is a Scientist within the Target Validation Group at LifeArc, located at the Stevenage Bioscience Catalyst science park. Victoria obtained her BSc in Biomedical Science, followed by an MRes in Biomedical Research and PhD in Molecular Cell Biology at Imperial College London. After completion
of her PhD in 2018, Victoria joined LifeArc where she is responsible for developing cellular assays for target validation and to support small molecule and antibody drug-discovery programmes.
References
- Grilo AL, Mantalaris A. (2019). The increasingly human and profitable monoclonal antibody market. Trends Biotechnol. 37, 9–16.
- Hoffmann RM, Coumbe BGT, Josephs DH, Mele S, et al. (2018). Antibody structure and engineering considerations for the design and function of Antibody Drug Conjugates (ADCs). OncoImmunology, 7(3), e1395127.
- Edmondson R, Broglie JJ, Adcock AF, Yang L. (2014). Three-dimensional cell culture systems and their applications in drug discovery and cell-based biosensors. Assay Drug Dev Technol. 12(4):207–218.
Related topics
Antibodies, Assays, Imaging, Oncology, T cells
Related conditions
Cancer
Related organisations
LifeArc
Related people
Laura Murch, Siobhán Leonard, Victoria Juskaite