Novel method for early disease detection using DNA droplets
Posted: 6 June 2022 | Ria Kakkad (Drug Target Review) | No comments yet
Researchers have developed a computational DNA droplet with the ability to recognise specific combinations of chemically synthesised microRNAs that act as biomarkers of tumours.


Aqueous droplet formation by liquid-liquid phase separation (or coacervation) in macromolecules is a popular topic in life sciences research. Of these various macromolecules that form droplets, DNA is most interesting as it is predictable and programmable, which are qualities useful in nanotechnology. Recently, the programmability of DNA was used to construct and regulate DNA droplets formed by coacervation of sequence designed DNAs.
Scientists at Tokyo University of Technology, Japan have developed a computational DNA droplet with the ability to recognise specific combinations of chemically synthesised microRNAs (miRNAs) that act as biomarkers of tumours. Using these miRNAs as molecular input, the droplets can give a DNA logic computing output through physical DNA droplet phase separation. Their findings were published in Advanced Functional Materials.
“The applications of DNA droplets have been reported in cell-inspired microcompartments. Even though biological systems regulate their functions by combining biosensing with molecular logical computation, no literature is available on integration of DNA droplet with molecular computing,” explained lead researcher, Professor Masahiro Takinoue.
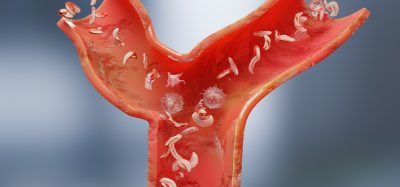
Developing this DNA droplet required a series of experiments. First, they designed three types of Y-shaped DNA nanostructures called Y-motifs A, B, and C with three sticky ends to make A, B, and C DNA droplets. Typically, similar droplets band together automatically while to join dissimilar droplets a special “linker” molecule required. So, they used linker molecules to join the A droplet with B and C droplet; these linker molecules were called AB and AC linkers, respectively.
In their first experiment they evaluated the “AND” operation in the AB droplet mixture by introducing two input DNAs. In this operation, the presence of input is recorded as one while its absence is recorded as 0. The phase separation of AB droplet mixture occurred only at (1,1), meaning when both input DNAs are present, suggesting successful application of AND operation. Following this study, the scientists decided to introduce breast cancer tumour markers, miRNA-1 and miRNA-2, to AC droplet mixture as inputs for the AND operation. The AND operation was successful implying that the computational DNA droplet identified the miRNAs.
In subsequent experiments, the team demonstrated simultaneous AND as well as NOT operations in AB mixture with miRNA-3 and miRNA-4 breast cancer biomarkers. Lastly, they created an ABC droplet mixture and introduced all the four breast cancer biomarkers to this solution. The phase separation in ABC droplet depended on the linker cleavage resulting in a two-phase separation or a three-phase separation.
This property of ABC droplet enabled the researchers to demonstrate the ability to detect a set of known cancer biomarkers or detect markers of three diseases simultaneously.
“If a DNA droplet can be developed which can integrate and process multiple inputs and outputs, we can use it in early disease detection as well as drug delivery systems. Our current study also acts as a steppingstone for research in developing intelligent artificial cells and molecular robots,” concluded Takinoue.
Related topics
DNA, Drug Development, microRNA, miRNAs, Nanotechnology, Technology
Related organisations
Tokyo University of Technology
Related people
Professor Masahiro Takinoue