Structural map of bacterial toxins identified
Posted: 18 September 2018 | Iqra Farooq (Drug Target Review) | No comments yet
A three-dimensional structural map of bacterial toxins has been identified for the bacterium Pseudomonas aeruginosa…

Researchers have mapped the three-dimensional structure of two toxic proteins that the bacteria, Pseudomonas aeruginosa, use to trigger the infection process.
Pseudomonas aeruginosa infections are common in hospitals. Those with reduced immunity, or large open wounds are susceptible to life-threatening infections by antibiotic resistant strains of the bacteria. This bacterium can also cause infections in the airways, making people with respiratory impairment vulnerable, and puts those with conditions like cystic fibrosis at risk.
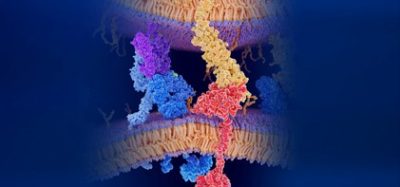
The researchers determined the structure of a human protein 14-3-3 (known to be necessary for the toxins to become active), and the toxins used to trigger the infection process, ExoS and ExoT. The team found how the protein acts as a ‘chaperone’, protecting the surface of the bacterial toxins through a large hydrophobic interaction interface. If the toxins are not protected, they form clusters in the cell’s water-soluble environment, and become inactive.
“Our study shows that it’s possible to block toxin activity with drug-like molecules via an interface where no other proteins have yet been shown to interact,” says principal investigator Dr Herwig Schüler, Associate Professor of Structural Biology at the Department of Biosciences and Nutrition, Karolinska Institutet.
The interface identified provides a possible target for drug compounds. The team identified two small molecules which could prevent the interaction between the protein and the toxins, and shows that toxins lose their toxic activity when these molecules are introduced.
Despite the effect being weak, it was observed and the researchers mention that it shows the principle works.
The previously known target site between the bacterial toxins and the human protein was also where other proteins in the cell bind. Thus, targeting this area would mean potentially serious side effects.
The researchers mention that the new site identified could be a more specific target.
The results of the study were published in Nature Communications.
Related topics
Disease Research, Drug Targets, Research & Development, Therapeutics
Related conditions
bacterial infections
Related organisations
Karolinska Institutet
Related people
Associate Professor Herwig Schüler