Sickle cell drugs could be tested with help from new computer model
Posted: 23 August 2019 | Rachael Harper (Drug Target Review) | No comments yet
New computer model that simulates the way red blood cells become misshapen by sickle cell disease could identity promising drug candidates.
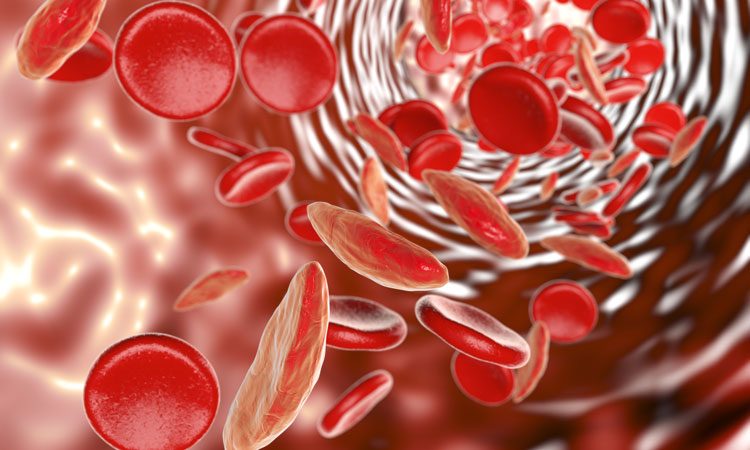

Researchers from Brown University have developed a new computer model that simulates the way red blood cells become misshapen by sickle cell disease which could be useful in the pre-clinical evaluation of drugs aimed at preventing the sickling process.
“We wanted to build a model that considers the entire sickling process and could be used to quickly and inexpensively pre-screen new drug candidates,” said Lu Lu, a PhD student in the Division of Applied Mathematics at Brown and the study’s co-lead author.
The model creates a single kinetic model of the entire sickling process. Using information gleaned from detailed supercomputer models, the researchers were able to build a simplified version that captures all the important dynamics of the sickling process, yet can be run on a laptop.
Because the dynamics of the sickling process can vary depending upon where in the body it’s happening, researchers designed the model to simulate sickling process in different organs. The model allows users to input parameters specific to the organ they are hoping to simulate. That same flexibility also enables to model to be run for individual patients who may have more or less severe versions of the disorder.
To test the potential effectiveness of drugs, the model allows users to input the mode of action by which a drug is presumed to work, information which is often gathered during preliminary lab studies.
“Sometimes a drug can be designed to work on one parameter, but ends up having a different effects on other parameters,” said George Karniadakis, a professor of applied mathematics at Brown and senior author of the new research.
The model can tell if those effects are synergistic or whether they may negate each other. So the model can give us an idea of the overall effect of the drug.”
The researchers are hopeful the model could be useful in identifying promising drug candidates.
“Clinical drug trials are very expensive and the vast majority of them are unsuccessful,” Karniadakis said. “The hope here is that we can do in silico trials to screen potential medications before proceeding to a clinical trial.”
The study was published in Science Advances.
Related topics
Drug Development, Informatics, Research & Development, Technology
Related conditions
Sickle cell disease (SCD)
Related organisations
Brown University, Science Advances
Related people
George Karniadakis, Lu Lu