Chemical markers that may unlock future therapeutic uses of messenger RNA
Posted: 28 April 2022 | Ria Kakkad (Drug Target Review) | No comments yet
Researchers have found new ways to artificially induce mRNA to respond in ways that could eventually lead to therapeutic outcomes.


A new study carried out by Hiroshima University’s Graduate School of Integrated Sciences for Life, Japan and Kansas State University, US aimed to find new ways to artificially induce messenger RNA (mRNA) to respond in ways that could eventually lead to therapeutic outcomes, opening up new possible genetic therapies. The researchers studied a biochemical process that adds a chemical mark to RNA bases, corresponding to a genetic letter of life’s blueprint, and identified such chemical marks that both speed up and slow down action in the beginnings of the chemical closures involved in generating gene-specified proteins. The study was recently published in Science Advances.
mRNA is called to action in the protein production process with a signal called the AUG Start Codon. The compound that AUG makes up is an amino acid called methionine, one of the 20 building blocks of protein molecules. Other RNA codons such as GUG (amino acid Valine), UUG (amino acid Leucine), and CUG (also Leucine) are generally considered “non-start” codons, meaning they are less likely to represent the beginning of a gene translation. Instead, they appear in the middle of protein coding region that is meant to open the genetic blueprint and produce a given protein.
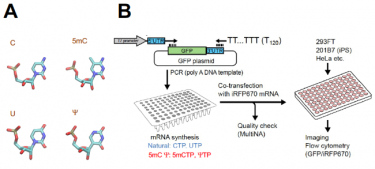

A, examples of chemically modified nucleotides used in this work shown to the right. Shown to the left are the natural nucleotide, cytosine (C) or uridine (U) before modification. B, experimental scheme. See legends to Fig. 1A-B of the original paper
[Credit: Katsura Asano, Hiroshima University and Kansas State University]
Few other codons than AUG are known to be able to activate mRNA in the way AUG does. But in setting out to change that, the team set out to test common RNA chemical modifications, evaluating their effects on different types of rare start codons initiating the translation process. To do so, they used their previous discovery that GUG, UUG, and CUG codons that are different by one letter from AUG, are converted to a reasonably strong start codon specifying methionine through attaching the optimum RNA sequence for initiating their translation event in animals. Their study design pitted a dozen RNA sequences, derived from these sequences, for expressing green fluorescent proteins through various non-AUG start codons at various efficiencies. To accurately evaluate GFP expression, they used a technique called flow cytometry to measure fluorescence from around 10,000 cells per attached RNA sequence and start codon. In this way, they compared translation efficiencies between natural RNA and chemically modified RNA.
They found common trends in altering translation efficiencies when a certain non-AUG start codon received a certain chemical mark. A remarkable discovery, they reported, was the ability of U-to-Psi (pseudouridine) conversion to dramatically increase initiation potentials of CUG, GUG and UUG start codons, and no effect on AUG.
The researchers hope the medical industry will take note of this new body of data and continue to conduct further research into how to use chemical modified RNA for generating synthetic expression switches, in such a way to stimulate translation activity in a highly targeted way in humans and animals.
“But there is a small chance that vaccines against retroviruses make vaccine cDNA when the patient encounters these viruses during immunisation. If this integrates into the patient’s genome, the antigen may be expressed in a way that attenuates vaccine production for boosting,” concluded Professor Katsura Asano, lead researcher. “But beyond that, the concept is so easy and adds no extra cost. So we hope these techniques are adopted.”
Related topics
Drug Targets, Protein, Research & Development, RNAs, Vaccine
Related organisations
Hiroshima University’s Graduate School of Integrated Sciences for Life, Kansas State University
Related people
Professor Katsura Asano