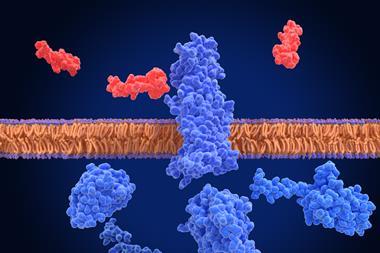
Researchers have designed synthetic, soluble versions of cell membrane proteins, which will enable faster and easier screening for new drugs.
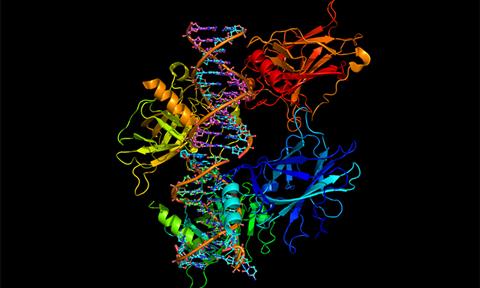
Researchers from the Laboratory of Protein Design and Immunoengineering (LPDI) at EPFL have used deep learning to design synthetic, soluble versions of cell membrane proteins that are frequently used in pharmaceutical research.
Numerous drug and antibody discovery pathways concentrate on folded cell membrane proteins. A chemical cascade that changes cellular behaviour is triggered when molecules of a drug candidate bind to these proteins. However, as these proteins are fixed in the lipid-containing outer layer of cells, they are difficult to access and are hydrophobic, making them challenging to study. Casper Goverde, a PhD student at LPDI commented: "We wanted to get these proteins out of the cell membrane, so we redesigned them as hyperstable, soluble analogues, which look like membrane proteins but are much easier to work with.”
The team designed a soluble protein analogue using their deep learning pipeline, and then used bacteria to produce modified proteins in bulk, which can bind directly in solution with molecular targets. This compares favourably to traditional screening methods, which are dependent on indirectly observing cellular reactions to drug and antibody candidates or extracting small quantities of membrane proteins from mammalian cells. “We estimate that producing a batch of soluble protein analogues using E. coli is around 10 times less expensive than using mammalian cells,” stated PhD student Nicolas Goldbach.
Recently, researchers had successfully utilised AI networks that use deep learning to design novel protein structures. However, in this study, the team required more accessible, soluble versions of proteins that already exist in nature. Goverde added: “We had the idea to invert this deep learning pipeline that predicts protein structure: if we input a structure, can it tell us the corresponding amino acid sequence?”
AlphaFold2, the structure prediction platform from Google DeepMind, was used to produce amino acid sequences for soluble versions of a variety of important cell membrane proteins, based on their 3D structure. The team then adopted a second deep learning network, ProteinMPNN, to optimise those sequences for functional, soluble proteins. Their method proved successful and accurate in producing soluble proteins that kept aspects of their native functionality, even when applied to very complex folds that have evaded other design methods thus far.
Notably, the pipeline successfully designed a soluble analogue of a protein shape called the G-protein coupled receptor (GPCR). A key pharmaceutical target, this represents around 40 percent of human cell membrane proteins. “We showed for the first time that we can redesign the GPCR shape as a stable soluble analogue. This has been a long-standing problem in biochemistry, because if you can make it soluble, you can screen for novel drugs much faster and more easily,” explained LPDI scientist Dr Martin Pacesa.
The results are proof-of-concept for the pipeline’s application to vaccine research, and cancer therapeutics.
This study was published in Nature.