Multi-omics approach yields novel insight for peanut allergies
Posted: 17 December 2019 | Drug Target Review | No comments yet
Researchers in New York have identified novel genes associated with the severity of peanut allergies and noted how these genes interact with other genes during allergic reactions. The findings could lead to better treatments for peanut allergies.
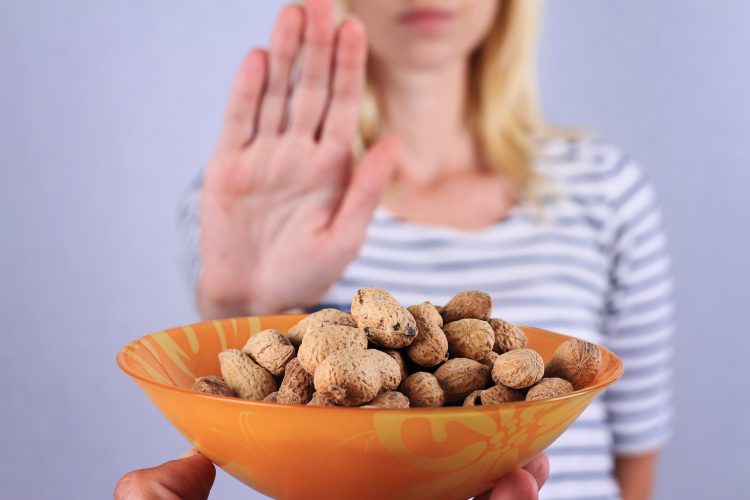


Peanut allergies are diverse in terms of levels of severity and are the leading cause of fatal food-related anaphylaxis. The tests used to determine the existence of a peanut allergy cannot distinguish between minor symptoms that might be experienced such as a rash or swelling, or life-threatening issues such as difficulty breathing or cardiovascular complications.
The study’s senior author, Supinda Bunyavanich, MD, MPH, Associate Professor of Genetics and Genomic Sciences, and Pediatrics, and Associate Director of the Elliot and Roslyn Jaffe Food Allergy Institute at Mount Sinai, was curious as to why the severity of reactions varies so much.
…[the researchers] also characterised ways in which these genes interact with other genes and CpG sites during allergic reactions”
To help address this question, Dr Bunyavanich and her team used novel multi-omic approaches to identify genes and networks of activity that might be driving the severity of peanut allergy reactions. The approaches included transcriptomics – the study of gene expression across the genome – and epigenomics – the study of reversible modifications to DNA that affect gene expression.
The study involved 21 children aged 7-17 with peanut allergies, who were given gradually increasing doses of peanut until they displayed an allergic response. The scientists sampled blood from the participants at three stages: before they ate, as they reacted, and after their reaction. The team confirmed their findings from the initial cohort by repeating the study in a further 19 children.
Taking blood samples at multiple times allowed the team to analyse both the transcriptome and epigenome (which can tell scientists which genes are turned on or off through a process called methylation) as the children reacted. Using this genome-wide approach, they identified more than 300 genes and 200 CpG sites (areas where DNA can be activated or inactivated by methylation) associated with reaction severity. Combining these data using integrative networks, the team also characterised key interactions between gene expression, CpG sites and reaction severity.
Having identified novel genes associated with the severity of peanut allergies, they also characterised ways in which these genes interact with other genes and CpG sites during allergic reactions to regulate biological processes. “It was very exciting to apply multi-omics to uncover how genes and methylation sites interact to affect reaction severity in these peanut-allergic kids,” says Anh Do, PhD, lead author of the study and postdoctoral fellow in the Bunyavanich Lab.
Important insights included that, while the findings support recognised roles for adaptive immunity in allergy, they also suggest that neutrophil (a type of white blood cell)-mediated immunity plays a prominent role in reaction severity. “We know neutrophil-mediated immunity is part of immune responses, and this study suggests it may play a central role in the severity of peanut allergic reactions,” said Dr Bunyavanich.
Additionally, one of the reaction severity drivers the team identified, the gene ARG1, can be inhibited by arginase inhibitors, a type of drug being researched for many disease applications. This study’s finding suggests it may also be a potential target for treating peanut allergies.
Dr Bunyavanich hopes future studies will identify biomarkers that can predict who is likely to have severe responses without having to expose them to peanuts first.
The findings were published in the Journal of Allergy and Clinical Immunology.
Related topics
Disease Research, DNA, Drug Leads, Drug Targets, Genetic Analysis, Targets
Related conditions
Peanut allergy
Related organisations
Icahn School of Medicine at Mount Sinai.
Related people
Anh Do, Supinda Bunyavanich