Researchers suggest IMPDH enzyme could be used to target sleeping sickness
Posted: 31 January 2020 | Hannah Balfour (Drug Target Review) | No comments yet
Research into drug targets for sleeping sickness parasite T. brucei has characterised the inosine-5′-monophosphate dehydrogenase (IMPDH) enzyme, providing a possible novel avenue for therapies.
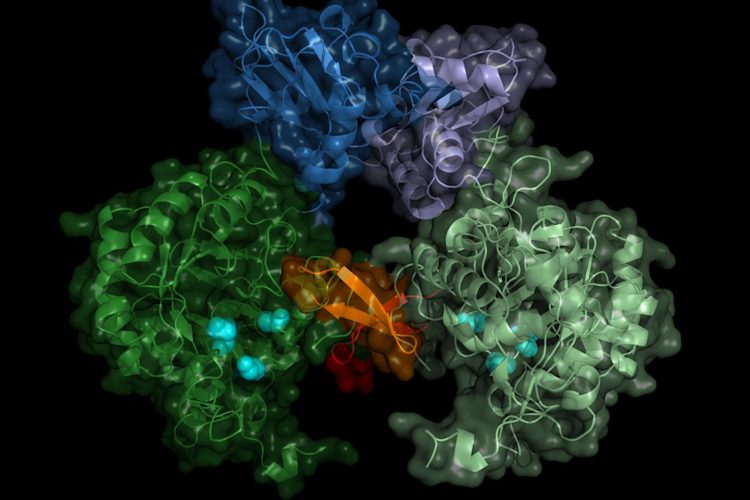

Structure of the parasite's IMP dehydrogenase. The active enzyme forms pairs (dimers), the "switch" region (Bateman region) is shown in shades of blue (credit: University of Lübeck/DESY, Lars Redecke).
Researchers using ‘in cellulo crystallisation’ and serial femtosecond crystallography have identified the precise structure of the sleeping sickness parasite’s inosine-5′-monophosphate dehydrogenase (IMPDH) enzyme. The team suggest their discovery could be used to produce highly effective selective and specific therapies against Trypanosoma brucei (T. brucei) in future.
…the data could open up pathways for inhibiting the T. brucei IMPDH in future”
Lars Redecke, co-author of the paper from the University of Lübeck and Deutsches Elektronen-Synchrotron (DESY), Germany, stated they focused on IMPDH because it “belongs to the central inventory of every [unicellular] organism and is an interesting target for drugs because it regulates the concentration of two vital nucleotides in the cell: guanosine diphosphate and guanosine triphosphate. The cell needs these nucleotides to supply energy and to build larger structures such as the genome. If you interrupt this cycle, the cell dies.”
In cellulo crystallisation, a method of creating crystals of a biomolecule (like enzymes) within the cells, was used by the team to freeze the structure of IMPDH molecules into tiny needles around 5 micrometres thick which protruded from the cells.
Bright X-rays were then used to analyse the tiny crystals: “X-ray lasers generate extremely intense flashes,” explained Henry Chapman, a Lead Scientist at DESY in the Center for Free-Electron Laser Science (CFEL). “Although the sensitive crystals evaporate immediately, they first generate a diffraction pattern from which the structure can be obtained.”
They recorded the diffraction patterns of approximately 22,000 microcrystals and calculated the structure of the enzyme, including what the researchers deemed the ‘on-off switch’ which can regulate IMPDH.
…researchers must now create an IMPDH inhibitor specific enough to block the parasite’s enzyme, but not the human version”
“The result does not only show the exact structure of the enzyme switch, the Bateman region, but also which molecules of the cell activate the switch and how these so-called co-factors bind to the enzyme switch,” revealed Karol Nass, co-author and PhD student who worked on the project at DESY.
The molecules which were observed to interact with the switch were adenosine triphosphate (ATP) and guanosine monophosphate (GMP). According to Christian Betzel from the University of Hamburg, a collaborator on the project, the data could open up pathways for inhibiting the T. brucei IMPDH in future: “One could think of constructing a kind of clasp that would cover the binding sites of both co-factors, for example.”
The researchers must now create an IMPDH inhibitor specific enough to block the parasite’s enzyme, but not the human version. Betzel indicated that, if successful, the method could potentially be extended to other pathogens.
He concluded: “Other parasites have a very similar structure and it might also be possible to attack those via the respective IMP dehydrogenase. The enzyme is a very interesting target for drugs, for example against the fox tapeworm or the elephantiasis pathogen.”
The paper was published in Nature Communications.
Related topics
Disease Research, Drug Targets, Enzymes, Imaging, Precision Medicine, Structural Biology
Related conditions
sleeping sickness
Related organisations
Deutsches Elektronen-Synchrotron (DESY), University of Lübeck
Related people
Christian Betzel, Henry Chapman, Karol Nass, Lars Redecke