Exploring gene interactions in influenza to improve accuracy of flu vaccines
Posted: 21 October 2022 | Ria Kakkad (Drug Target Review) | No comments yet
Researchers from the University of Illinois at Urbana-Champaign found the evolutionary potential of influenza A virus haemagglutinin is extremely restricted by epistatic interactions with neuraminidase.

The influenza virus, which causes the flu, is a major public health issue, infecting millions of people and is estimated to cost $10 billion in direct medical costs in the US annually. Like most viruses, influenza mutates rapidly as it spreads, making it difficult to vaccinate against every possible strain. As a result, every year there is a massive effort to determine which strains will likely be the most prevalent, to make a vaccine that offers the best protection for that season.
“It is critical to better understand the fundamental rules that govern the evolution of viruses and how they escape from our immune systems,” said Associate Professor Chris Brooke, from University of Illinois Urbana-Champaign, US.
Researching flu vaccines
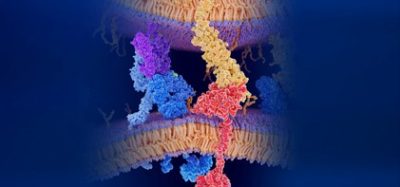
Most research efforts for influenza vaccination focus on the evolutionary potential of the surface protein haemagglutinin (HA) because this is the main protein the immune system targets. HA binds to receptors on the surface of cells, allowing the influenza virus to enter and replicate. However, another surface protein called neuraminidase (NA) has been largely overlooked. NA is important for later in the virus life cycle, when it destroys the receptors originally used for entry, cleaving the cell and releasing the virions inside. HA and NA are highly functionally involved with each other, despite their opposing actions, because they are both necessary for the virus to infect cells and spread.
In a new paper published in Cell Host & Microbe, researchers from the University of Illinois at Urbana-Champaign explored how changes in NA activity affected the evolutionary potential of HA. To do this, they used a wildtype influenza strain and two strains identical to the wildtype except for a mutation that reduced NA activity. Next, the team used a process called deep mutational scanning to create a library of HA mutated versions of their three strains of influenza. The researchers could then measure HA activity and fitness across the mutated virus strains.
It is critical to better understand the fundamental rules that govern the evolution of viruses and how they escape from our immune systems”
“This is a high-throughput method for introducing every possible amino acid substitution into a given region of interest and then measuring the effects of those substitutions on relative fitness,” said Brooke. “We can then quantify fitness effects of specific substitutions depending on the NA gene they were paired with.”
How does HA interact with NA?
Through this process, the team determined that virus strains with reduced NA activity had higher mutational tolerance in changes in HA. In other words, when the influenza strains with lower NA receptor binding were paired with mutations that also decreased HA receptor binding, they largely showed no decrease in fitness, compared to the wildtype strain, which often did have decreased fitness. The researchers explained that this is likely due to the opposing functions of HA and NA, where the former works to break into the cell, and the latter works to escape the cell.
“If NA has a substitution that decreases its activity, we see compensatory substitutions in HA that lower its relative activity as well, bringing them back into balance. If only one of them goes up or down, the cell becomes out of balance and this decreases the overall fitness. So, you will see a compensatory substitution to bring it back into the optimal range,” explained Brook.
“I think we were all surprised to see that,” said Tongyu Liu, a graduate student in Brooke’s lab and lead author on the paper. “The textbook view of mutations is that they are mostly deleterious. But here in our experiment, we demonstrated that the interaction between HA and NA could reshape the fitness effect of mutations, from mostly deleterious to primarily neutral.”
LONG READS: Is an HIV vaccine on the horizon?
READ HERE
The researchers also grew the same strains with reduced NA activity in the presence of neutralising antibodies that target HA to see what mutations would arise. The experiment mimics what happens in the body, when immune cells target and attack the HA protein of influenza cells to clear an infection. By looking at what HA variants are selected to evolve in this environment, the researchers found more evidence that the pathways that HA evolution takes to escape immune pressure are highly contingent on NA function.
The evolution of HA is mainly studied in isolation from other genes in influenza vaccine research, but Brooke said this data demonstrates the need to look at the interactions between HA and genes for other proteins, like NA, to predict better how HA will evolve depending on mutations in these other genes across strains.
The future of flu vaccine research
“Every year we try and pick which of the many different co-circulating flu strains to target with the next year’s vaccine and if we do not pick correctly, that leads to more severe infections and deaths,” Brooke said. “So it is really important to understand the rules that govern the evolution of the virus so that we can better predict the specific pathways it will take.”
The researchers now hope that their findings will be used to improve accuracy for predicting future predominant genotypes and create vaccines against these strains.
Related topics
Genetic Analysis, Immunology, Molecular Targets, Vaccine
Related conditions
Influenza A
Related organisations
Carl R Woese Institute for Genomic Biology University of Illinois At Urbana-Champaign, University of Illinois at Urbana-Champaign
Related people
Associate Professor Chris Brooke, Tongyu Liu