What are the pros and cons of using organoids?
Posted: 30 August 2019 | Dr Shona Lang | No comments yet
Dr Shona Lang investigates the advantages and disadvantages of using organoids within R&D, highlighting the most important questions to ask before using these models.
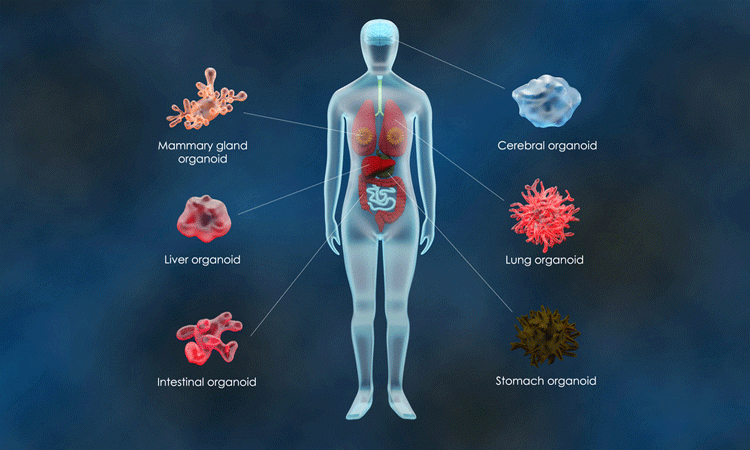
Organoids are three-dimensional (3D) cell cultures which mimic tissue architecture. Their growing importance in various fields of research saw them named ‘Method of the Year 2017’1 and their use in drug testing has seen a large increase in the market for 3D cell cultures.2 Importantly, these models have revealed the ability of the cell to differentiate and self-organise. Researchers have found organoids relatively easy to grow and they can produce stunning images; they can be observed using molecular, cellular and imaging techniques.
However, like any other model derived from patient tissues, they must be clearly validated. So, if you are thinking of investing in an organoid model or you are trying to select the best model from several possibilities, what questions should be asked? There are clear advantages in using organoids, but often the negatives can be overlooked or are simply unpublished.
Not an average culture
Organoids are an upgrade from the traditional primary cultures grown as a monolayer in a tissue culture flask. The difference for organoids is that cells are grown within a basement membrane gel and develop into 3D shapes; primarily into hollow or budding spheres. The cells seeded into this type of culture system can be selected (eg, stem cells) or unselected to reflect the cellular heterogeneity of the patient.
If you identify an organoid with a mutation of interest, you need to confirm what proportion of cells have that mutation”
Although organoids are now widely used, they are also under-investigated and often poorly validated. It is not known what effect the basement membrane gel has on cellular behaviour – does it support the natural differentiation of the cell or does it reprogramme growth in an undetermined way? This must be kept in mind if you assume that your model recreates the tissue of interest.
Organoid morphology
Organoids offer superior morphology if you are studying a glandular tissue, but they are not appropriate for studying stratified tissues, such as skin. Originally organoid cultures were grown to investigate normal cellular differentiation in the prostate3 and breast.4 Their use as a tumour model is debatable as the influence of the basement membrane gel is not currently understood. One of the classical definitions of tumour growth is anchorage independence, yet organoids are anchorage dependent due to their adherence to the basement membrane proteins in the gels. In addition, tumour cells known to grow as solid masses in vivo can grow as hollow spheres5 in the organoid system, so they do not always recreate patient tissue architecture. Therefore, it is important to carefully consider the morphology of the disease you are studying.
Purity and removing contaminants
Epithelial cells.
Cultures derived from tissues will contain a mix of cell types, for example, epithelial or stromal. These can be contaminated by neighbouring tissues dependent on the sampling technique, eg, prostate tissue sampled by transurethral resection could contain urethral tissue. Tumour tissues can also be contaminated by normal cells. A study of in vitro and in vivo models for precision medicine indicated that between 25-95 percent of ‘tumour’ organoids (across a range of tissues) were normal cells.6 When using organoids, it is imperative to confirm the absence of all potential contaminants.
Heterogeneity of organoids
Although heterogeneity is frequently reported as a positive aspect of spheroid cultures, it must also be recognised as a negative. If you are testing a drug and want to know its effect on a heterogeneous culture of cells, then this is a positive characteristic. However, when investigating the action mechanisms of a drug and which cells are targeted, using a heterogeneous mix of cells will not yield answers. Evidence suggests that you cannot rely on the in situ genotype to reflect the in vitro genotype.7 Since cells are usually seeded at high density into an organoid culture there is also a risk that a single organoid may not be clonal (derived from a single cell). If you are using a heterogeneous mix of cells it is critical to check whether the ratio and type of cells (tumour, normal, different tumour clones) reflects the patient or whether it is better to use multiple samples or organoid cultures per patient.
Reproducibility and clonal drift
Although organoids are now widely used, they are also under-investigated and often poorly validated”
Heterogeneous cultures contain many different clones, which means that as the culture grows certain clones will dominate while others will die out. Therefore, the heterogeneity will be different from one pass to the next and experimental reproducibility will suffer.8 Whilst a mutation may be detectable in a culture of organoids this does not mean that 100 percent of the cells within or between organoids are tumourigenic and this will have consequences for drug testing. If you identify an organoid with a mutation of interest, you need to confirm what proportion of cells have that mutation. If 10 percent of the cells contain the mutation, the response to the drug will be dominated by the wild type cells. This lack of validation can lead to false negatives and poor reproducibility. Accordingly, understanding the genotype and phenotype of your culture (including the proportion of cells) at every passage to ensure your model is fit for application to precision medicine is important.
Evidence-based validation
Currently there is a growth in omics research – the study of large datasets derived from genomic and proteomic research – that has enabled an understanding of the complete molecular profile of the patient and helped establish personalised medicine. Omics research has also led to the establishment of numerous databases to store information, which can be used to source and categorise research material, eg, biobanks and cancer model directories. However, the validity of the data in these databases is often unclear and currently there are calls to improve research standards9 and rigour10 in this area. It is important to remember that the identification of a gene mutation within a model does not confirm that the mutation is present in all cells or that it will last over a period in culture.
 Currently the validation of pre-clinical models is based on the evidence reported in scientific publications or the opinion of experts; there are biases in both.11 With the advance of the Open Science movement it is well known that scientific publications face problems with publication bias.11 Expert opinion is open to bias, conflicts of interest and spin.12 Evidence-based decision making has been used for several years in medicine to avoid partiality.13 This process relies on developing a research question and protocol to find and evaluate all available evidence. It is advantageous because not only does it evaluate evidence in terms of quality, but it also highlights the evidence gaps. Advances in the application of evidence-based decision making for biomedical science allows investors and researchers to find the best validated models and ensure they recapitulate the disease of interest.14 Decision-making tools allow researchers to identify which validation criteria have been met and those which have not (tissue of origin proven, cell lineage proven, contaminants absent, concordance of histology and genotype, concordance of response to drug treatment). This ensures that both useful evidence and evidence gaps can be identified.15 The process aids the selection of the best models and can highlight the need for further validation experiments.
Currently the validation of pre-clinical models is based on the evidence reported in scientific publications or the opinion of experts; there are biases in both.11 With the advance of the Open Science movement it is well known that scientific publications face problems with publication bias.11 Expert opinion is open to bias, conflicts of interest and spin.12 Evidence-based decision making has been used for several years in medicine to avoid partiality.13 This process relies on developing a research question and protocol to find and evaluate all available evidence. It is advantageous because not only does it evaluate evidence in terms of quality, but it also highlights the evidence gaps. Advances in the application of evidence-based decision making for biomedical science allows investors and researchers to find the best validated models and ensure they recapitulate the disease of interest.14 Decision-making tools allow researchers to identify which validation criteria have been met and those which have not (tissue of origin proven, cell lineage proven, contaminants absent, concordance of histology and genotype, concordance of response to drug treatment). This ensures that both useful evidence and evidence gaps can be identified.15 The process aids the selection of the best models and can highlight the need for further validation experiments.
In conclusion, evidence-based decision-making tools improve translation into the clinic by reminding investors of the important questions to ask. It is vital to always be wary of models with minimal validation.
About the author
Dr Shona Lang has spent eight years working in clinical epidemiology preparing systematic reviews, meta-analyses, rapid reviews and health technology assessments to support policy making, licensing and reimbursement decisions in clinical research. She has established QED Biomedical Ltd and is researching the value of evidence-based decision making in biomedical science and clinical translation and hopes to provide solutions to de-risk investment in this field.
References
- Method of the Year 2017: Organoids [Internet]. Nature Methods. 2018 [cited Aug 2019]. 15(1):1–1. Available from: http://www.nature.com/articles/nmeth.4575
- Sullivan L. 3D Cell Culture: Global Markets and Out-of-this-World Applications Research [Internet]. BBC Research. 2017 [cited Aug 2019]. Available from: https://blog.bccresearch.com/3d-cell-culture-global-markets-and-out-of-this-world-applications
- Lang SH, Stark M, Collins A, Paul AB, Stower MJ, Maitland NJ. Experimental Prostate Epithelial Morphogenesis in Response to Stroma and Three-Dimensional Matrigel Culture [Internet]. Cell Growth Differ. 2001 [cited Aug 2019]. 1;12(12):631–40. Available from: http://cgd.aacrjournals.org/cgi/content/abstract/12/12/631
- Barcellos-Hoff MH, Aggeler J, Ram TG, Bissell MJ. Functional differentiation and alveolar morphogenesis of primary mammary cultures on reconstituted basement membrane [Internet]. PubMed – NCBI. 1989 [cited Aug 2019]. 105(2):223–35. Available from: https://www.ncbi.nlm.nih.gov/pubmed/2806122
- Lang SH, Sharrard RM, Stark M, Villette JM, Maitland NJ. Prostate epithelial cell lines form spheroids with evidence of glandular differentiation in three-dimensional Matrigel cultures. [Internet]. British Journal of Cancer. 2001 [cited Sept 2018]. 85(4):590–9. Available from: http://www.nature.com/doifinder/10.1054/bjoc.2001.1967
- Pauli C, Hopkins BD, Prandi D, Shaw R, Fedrizzi T, Sboner A, et al. Personalized In Vitro and In Vivo Cancer Models to Guide Precision Medicine. [Internet]. Cancer Discovery. 2017 [cited Aug 2019]. 7(5):462–77. Available from: http://cancerdiscovery.aacrjournals.org/lookup/doi/10.1158/2159-8290.CD-16-1154
- Ozbun M, Medina D, Butel J. p53 mutations in mouse mammary epithelial cells: instability in culture and discordant selection of mutations in vitro versus in vivo. [Internet]. Cell Growth Differ. 1993 [cited Aug 2019]. 1;4(10):811–9. Available from: http://cgd.aacrjournals.org/cgi/content/abstract/4/10/811
- Huch M, Knoblich JA, Lutolf MP, Martinez-Arias A. The hope and the hype of organoid research. [Internet]. Development. 2017 [cited Aug 2019]. 144(6):938–41. Available from: http://dev.biologists.org/lookup/doi/10.1242/dev.150201
- Volkmann A, De Bin R, Sauerbrei W, Boulesteix A-L. A plea for taking all available clinical information into account when assessing the predictive value of omics data. [Internet]. BMC Medical Research Methodology. 2019 [cited Aug 2019 Aug]. 19(1). Available from: https://bmcmedresmethodol.biomedcentral.com/articles/10.1186/s12874-019-0802-0
- Riley P. Three pitfalls to avoid in machine learning. [Internet]. Nature. 2019 [cited Aug 2019]. 572(7767):27–9. Available from: http://www.nature.com/articles/d41586-019-02307-y
- Ioannidis JPA. Why Most Published Research Findings Are False. [Internet]. PLoS Medicine. 2005 [cited Jan 2019] 2(8):e124. Available from: https://dx.plos.org/10.1371/journal.pmed.0020124
- Boutron I, Haneef R, Yavchitz A, Baron G, Novack J, Oransky I, et al. Three randomized controlled trials evaluating the impact of “spin” in health news stories reporting studies of pharmacologic treatments on patients’/caregivers’ interpretation of treatment benefit. [Internet]. BMC Medicine. 2019 [cited Aug 2019]. 17(1). Available from: https://bmcmedicine.biomedcentral.com/articles/10.1186/s12916-019-1330-9
- Sackett DL, Rosenberg WMC, Gray JAM, Haynes RB, Richardson WS. Evidence based medicine: what it is and what it isn’t. [Internet]. BMJ. 1996 [cited Aug 2019]. 312(7023):71–2. Available from: http://www.bmj.com/cgi/doi/10.1136/bmj.312.7023.71
- Collins A, Ross J, Lang SH. A systematic review of the asymmetric inheritance of cellular organelles in eukaryotes: A critique of basic science validity and imprecision. PLOS ONE. 2017;12.
- Collins AT, Lang SH. A systematic review of the validity of patient derived xenograft (PDX) models: the implications for translational research and personalised medicine. [Internet]. PeerJ. 2018 [cited Jan 2019]. 6:e5981. Available from: https://peerj.com/articles/5981
Related topics
Drug Development, Drug Targets, Genomics, Organoids, Research & Development, Stem Cells