A rational approach to discover and precision-engineer new biologic drugs
Posted: 2 December 2020 | Jerome Boyd-Kirkup (Hummingbird Bioscience) | No comments yet
When it comes to developing antibody drugs, Dr Jerome Boyd-Kirkup and his team are not sticking to the classical playbook. Here, he explains how they use systems biology and immuno-engineering to remove the element of luck from biologic drug discovery and development.
THE DEVELOPMENT of biologics, particularly antibodies, for therapeutic applications is notoriously risky. Unlike the development of small molecules, biologics do not benefit from decades of experience to guide decision-making. Questions remain, such as which biological disease mechanisms are amenable to therapeutic intervention? How can we develop molecules that could deliver such intervention? What is the best way to test and manufacture these molecules to obtain regulatory approval and enable clinical use? The fresh unknowns, driven by the involvement of living cells and organisms, introduce new variables, stochastic variation and a lot of empiricism, which impinge on the success of traditional approaches to biologic drug development.1,2
Despite these challenges, biologics have transformed medicine, providing many patients with new drugs for diseases that were previously untreatable. Seven of the top 10 best-selling drugs worldwide are currently biologics.3 Given their enormous success and the promise of additional benefits to multiple conditions, it is critical to find new and more efficient ways of discovering and developing these drugs. We believe that by applying systems biology and immuno-engineering, we can take a more precise approach. To give a tangible example, we explain here how our unique Rational Antibody Discovery (RAD) platform is being used to develop novel antibody drugs.
Developing a novel drug for cancer
Despite it being a highly promising target, HER3 has proved difficult to drug effectively”
Our investigational drug, HMBD-001, was developed to treat tumours that have aberrant activation of the cancer-associated protein, HER3. A member of the human epidermal growth factor receptor (HER/EGFR) family, HER3 is known to drive cancer cell growth through dimerisation with HER2 or EGFR. Dimerisation is favoured in the presence of the ligand, neuregulin 1 (NRG1) or HER2/EGFR over-expression. In recent years, HER3 has been implicated in a significant number of cancers, including breast, gastric, pancreatic and lung cancers,4-6 and in acquired resistance to other therapies.7 Moreover, a subpopulation of cancer patients has been identified with NRG1 fusions; that is, the hybridisation of the NRG1 gene with another gene to produce NRG1 fusion proteins that potently activate HER3.6
Despite it being a highly promising target, HER3 has proved difficult to drug effectively. Attempts by other drug developers to target HER3 with antibodies that block NRG1 binding have thus far yielded suboptimal results in the clinic. We postulated that this was likely due to the inability of these antibodies to completely prevent dimerisation of HER3, since dimerisation is not fully dependent on NRG1 binding.
We were able to solve this by rationally discovering and engineering a therapeutic antibody that would successfully block all HER3 dimerisation, irrespective of NRG1 binding. The key to this was the RAD platform (see Figure 1).
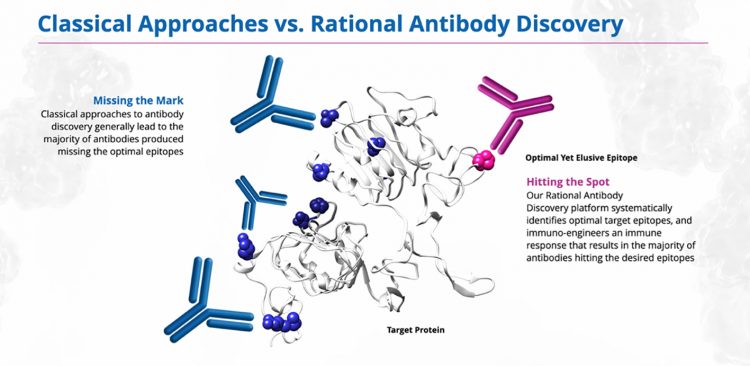
Figure 1: The Rational Antibody Discovery (RAD) platform has the potential to revolutionise approaches to drug discovery
A pioneering platform
Our platform is comprised of two key aspects: mAbPredict and mAbHits (see Figure 2). mAbPredict relies on computational biology to integrate vast amounts of data from multidisciplinary sources with the help of machine-learning algorithms. The data is then interrogated to firstly ensure that the treatment target, in this case HER3, is strongly validated in terms of its critical role in the disease mechanism and its viability as a drug target; and secondly, to make predictions regarding which regions of the protein should be targeted for optimal antibody efficacy and safety profiles.
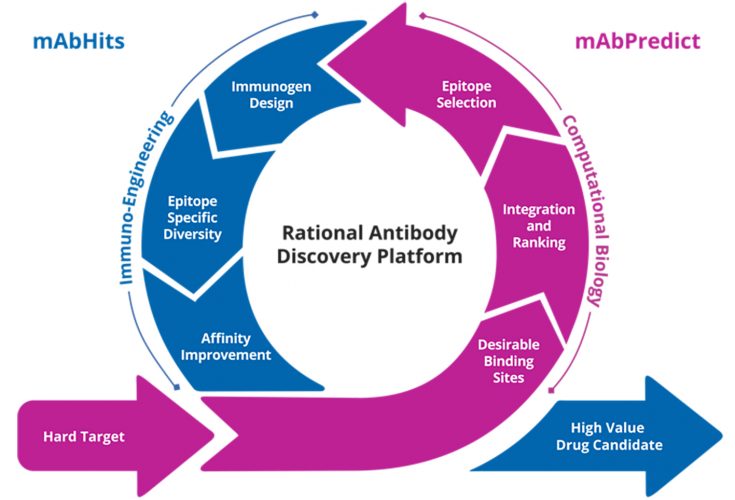
Figure 2: The Rational Antibody Discovery (RAD) platform applies computational biology and immuno-engineering to discover and develop high value drug candidates
In our investigation, we found that the optimal epitope on HER3 was located in the domain of the protein that interfaces with HER2 and EGFR during dimerisation and, importantly, was available for antibody binding in both closed and open conformations of the protein. Binding this epitope was predicted to directly interfere with the ability of HER3 to dimerise and thus prevent its activation, regardless of ligand binding. This optimal epitope is not represented in any of the previous HER3 antibodies from other drug developers, which tend to bind to epitopes in the ligand-binding domain of the protein (see Figure 3).
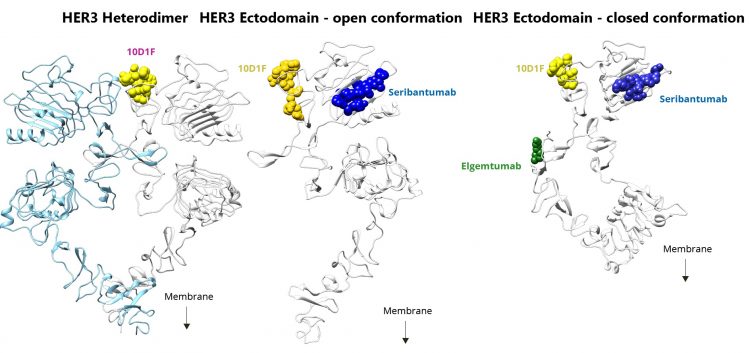
Figure 3: The antibody, HMBD-001, uniquely binds to an epitope that directly blocks HER3 dimerisation in both open and closed conformations
To circumvent this and shift the production of antibodies towards epitopes that are not immuno-dominant, we precisely modulate the immune system of the antibody-producing organism using mAbHits, our proprietary immuno-engineering platform. This involves careful design and delivery of a range of immunogens and timed pharmacologic manipulation of the underlying mechanisms driving the immune response, in order to develop a directed B-cell response to the epitope in the disease-associated conformation of the protein. The result is a diverse range of antibodies that target the optimal yet elusive epitope, providing many options from which to select a lead drug candidate. Although we may further engineer the antibodies if necessary, the affinity of the antibodies we discover is often very high as the in vivo mechanisms of B cell maturation are leveraged to mature optimal antibodies prior to isolation. This was how HMBD-001 was discovered and developed. One of the reasons for this is that conventional antibody discovery is generally unsupervised, relying on chance to deliver antibodies that satisfy design goals for therapeutics. Whether by immunisation methods or display methods, the hurdle is considered to be a screening challenge such that if you can sample the antibody repertoire deeply enough you will discover the most potent antibody that binds the optimal epitope. However, the reality is that the germline antibody repertoires reflected in display methods do not bind to all regions of a protein, nor does the immune system select and mature antibodies against all regions of a protein during an immunisation campaign. Antibodies tend to cluster around “immunodominant” regions of a protein and it is purely chance if these regions also coincide with regions where binding causes therapeutic modulation of the protein function.
From concept to clinical care
A large body of pre-clinical data from cell and animal models has demonstrated that HMBD‑001 potently inhibits the activation of HER3 and its signalling pathway, independent of any NRG1 binding, leading to dramatic inhibition of tumour growth.4,7 Thus, HMBD-001 represents a unique, highly specific, anti-HER3 neutralising antibody with a novel mechanism of action against HER3‑activated tumours. HMBD-001 is now nearing clinical trials in partnership with Cancer Research UK (CRUK) and we anticipate that it will be the best-in-class for treating HER3-activated cancers, including those with NRG1 fusions.
Our immuno-engineering capability also enables us to overcome tolerance to species-conserved
epitopes and generate targeted agonist antibodies”
HMBD-001 showcases the advantages of our approach and platform in designing functional antibodies. However, the RAD platform can also address other key challenges encountered in classical methods of drug discovery and development. For example, we have applied the platform to address epitope instability, as is seen in viruses, by predicting the epitopes most likely to be conserved in rapidly mutating proteins and immuno-engineering an antibody response against these conserved epitopes. Our immuno-engineering capability also enables us to overcome tolerance to species-conserved epitopes and to generate targeted agonist antibodies, which must bind to a very specific region of the target protein to achieve potent activation. Moreover, we have successfully drugged multipass transmembrane proteins, such as G-protein-coupled receptors (GPCRs) and ion channels, by overcoming the competition from other immunogenic epitopes to generate antibodies that bind to these membrane-bound proteins in their native conformation.
In a relatively short space of time, we have used the platform and in-house capabilities to build a rich pipeline of first- and best-in-class drug candidates in oncology, autoimmune and infectious diseases, together with global partners in academia and industry. Aside from HMBD-001, our other lead programmes include an anti-VISTA antibody, HMBD-002, and an anti‑SARS-CoV-2 antibody, HMBD-115 – one of the first COVID-19 antibodies in the world to enter clinical trials.
We strongly believe that the latest advances in systems biology and data science can help us better understand and solve the underlying causes of disease. By taking a rational approach, we aim to overcome the challenges of classical methods of therapeutics discovery and profoundly improve the way we engineer revolutionary therapies for hard-to-drug targets to deliver a new generation of transformative medicines to patients.
About the author
Dr Jerome Boyd-Kirkup is the Co-Founder and Chief Scientific Officer of Hummingbird Bioscience. He is responsible for overseeing the development of new technologies and intellectual property, designing experimental strategies for drug discovery, leading the development and testing of novel therapeutics and overseeing the drug pipeline. He is a molecular systems biologist by background with extensive experience at the interface of computational and wet lab biology and a career research focus of applying systems biology approaches to tackling human disease. Before co-founding Hummingbird Bioscience in 2014, he was an academic at institutions in the UK and China.
References
- Morrow T, Felcone LH. Defining the difference: What makes biologics unique. Biotechnol Healthc. 2004 Sep; 1(4): 24–29.
- Challener CA, Branch E, Tiene G. Challenges for next-generation biological therapeutics in discovery and development. Pharma’s Almanac. May 29, 2018. Available from: https://www.pharmasalmanac.com/articles/challenges-for-nextgeneration-biological-therapeutics-discovery-and-development (accessed Sep 2020).
- Analysis Report. The top selling prescription drugs by revenue. Pharmaceutical Technology. May 21, 2019 – updated Jan 27, 2020. Available from: https://www.pharmaceutical-technology.com/features/top-selling-prescription-drugs/ (accessed Sep 2020).
- Boyd-Kirkup J, Thakkar D, Ingram PJ. HMBD001, a novel anti-ErbB3 antibody with a unique mechanism of action, effectively inhibits tumor growth in pre-clinical models of ErbB3+ solid tumors. Cancer Research, 77(13 Supplement), 24–24. Available from: https://doi.org/10.1158/1538-7445.AM2017-24 (accessed Sep 2020).
- Baselga J, Swain SM. Novel anticancer targets: revisiting ERBB2 and discovering ERBB3. Nat Rev Cancer. 2009; 9: 463–475.
- Hynes NE, MacDonald G. ErbB receptors and signaling pathways in cancer. Curr Opin Cell Biol. 2009; 21: 177–184.
- Thakkar D, Sancenon V, Taguiam M, Guan S, Wu Z, Ng E, Paszkiewicz KH, Ingram PJ, Boyd-Kirkup JD. 10D1F, an anti-HER3 antibody that uniquely blocks the receptor heterodimerization interface, potently inhibits tumor growth across a broad panel of tumor models. Mol Cancer Ther. 2020; 19(2): 490–501.
Related topics
Biologics, Biopharmaceuticals, Drug Discovery, Drug Leads, Hit-to-Lead, Oncology
Related conditions
Breast cancer, Cancer, Covid-19, Gastric cancer, Lung cancer, Pancreatic cancer