Pacemaker channel discovery could lead to better heart drugs
Posted: 11 November 2022 | Ria Kakkad (Drug Target Review) | No comments yet
The research has shown in high-resolution detail how certain lipids interact with pacemaker ion channels to enhance their activity.

In a new study, researchers from Weill Cornell Medicine, US, used advanced methods including cryogenic electron microscopy (cryo-EM) to show in high-resolution detail how certain lipids interact with pacemaker ion channels to enhance their activity. In principle, modulating this lipid interaction with a drug could be a good strategy for treating cardiac arrhythmias and other conditions. The study was recently publishing in Nature Structural & Molecular Biology.
“Ion channels have been notoriously difficult to target with drugs, because of the challenge of identifying drug binding sites that are specific to particular channels, but this work reveals sites that might be highly specific and thus viable drug targets,” said senior author Dr Crina Nimigean.
Ion channels are tube-like protein structures that reside within the membranes of a cells, enabling and regulating flows of charged molecules of potassium, sodium and other electrolytes into and out of the cell. These flows of charged molecules, or ions, are key determinants of cell behaviour. Specialised ion channels called pacemaker channels are particularly important for the rhythmic activities of heart cells and neurons. Understanding precisely how these pacemaker channels work could thus lead to better treatments for cardiac arrhythmias, chronic pain, epilepsy and other conditions. Arrhythmias alone are estimated to affect millions of people in the United States.
Scientists have known that lipids, the major constituents of cell membranes, are involved in modulating the activities of pacemaker channels. But they have not known much about how these interactions work. In general, methods for studying lipids have been less well developed than methods for studying proteins and other biological molecules. Moreover, ion channels function while embedded in the cell membrane, which is made of two thin layers of lipids—and that membrane and its lipid components have been hard to replicate in a way that allows detailed experimental manipulation.
The team were able to obtain important clues about lipid-pacemaker channel interactions by first examining a bacterial pacemaker channel, SthK. In prior studies, they had developed an experimental platform for studying SthK, including a membrane-like environment that they could alter experimentally. SthK is a useful model for pacemaker channels found in humans—known as HCN channels—because the two channels, despite the evolutionary gulf between them, have many important similarities, they said.
The SthK studies revealed that two lipids, phosphatidyl-glycerol and cardiolipin, can bind to the channel in a way that disrupts a specific molecular connection, known as a salt bridge, which normally tends to clamp the channel shut. When the salt bridge was disrupted by the lipids, the channel became more open and active. Similarly, removing the salt bridge by other means dramatically increased channel activity and abolished the ability of these lipids to affect that activity.
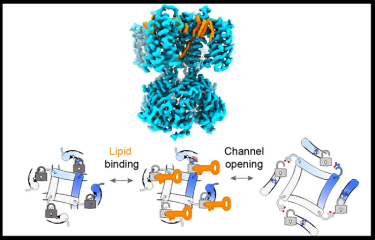

Lipids bind to pacemaker ion channels to unlock their gates. Top: Side view of the structure of SthK channels (blue) with bound lipids (orange). Bottom: Bottom view of the channel pore opening as it transitions, from left to right, from closed and locked, to closed and unlocked, to open. Channel protein is in blue and grey cartoon, lipids are represented by orange keys, channel locked closed is indicated by a dark grey lock, channel unlocked is indicated by a light grey lock, and channel open is indicated by the enlargement of the pore size (rightmost panel). The lipid binding and channel opening transitions are indicated above the arrows
[Credit: Weill Cornell Medicine].
The same salt bridge is found in HCN channels, and experiments suggested that in the latter the same lipid-modulation mechanism is at work: Removing the salt bridge had the same effect on channel activity as in the SthK experiments, although for HCN channels the key lipid binding partner was a different lipid, phosphatidic acid.
The experiments included cryo-EM imaging of lipids binding to SthK—imaging that offers clues to how a future drug might disrupt or enhance this lipid interaction to modulate pacemaker channel function.
The researchers hope that in follow-on work they can illuminate the role of this lipid-pacemaker interaction in abnormal conditions such as arrhythmias or cancers.
“Various conditions can affect the lipid composition of the heart and other tissues, so it would not be surprising to see this pacemaker mechanism altered in diseases,” concluded first author Dr Philipp Schmidpeter.
Related topics
Drug Development, Drug Discovery, Lipidomics, Lipids, Microscopy
Related conditions
cardiac arrhythmias
Related organisations
Weill Cornell Medicine
Related people
Dr Crina Nimigean, Dr Philipp Schmidpeter