CRISPR-Cas9 gene editing imaged for first time
Posted: 9 July 2019 | Victoria Rees (Drug Target Review) | No comments yet
Researchers have generated images of CRISPR-Cas9 gene editing for the first time, enabling improvement of the technique.
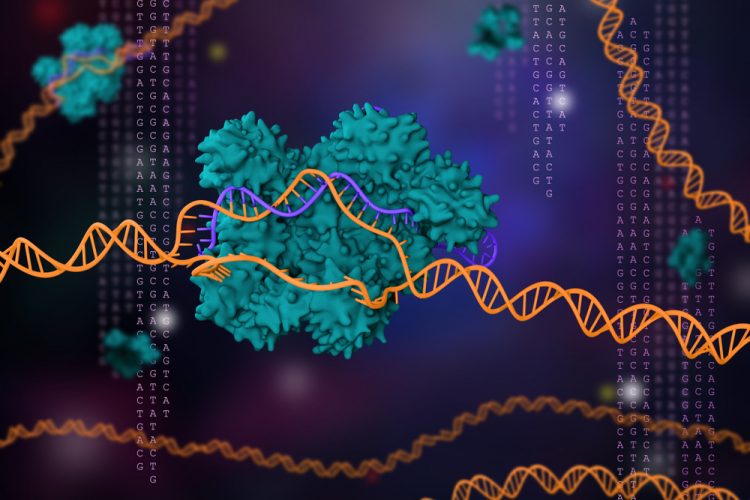
For the first time, researchers have been able to capture high-resolution, three-dimensional images of an enzyme in the process of editing DNA strands. According to the team, their findings will aid in developing gene editing techniques that operate more efficiently and precisely.
The team, from the University of British Columbia (UBC), used cryogenic electron microscopy (cryo-EM) to reveal the workings of CRISPR-Cas9. The unprecedented images provide a glimpse of the molecular motions occurring in the course of DNA cutting.
“These images provide us with invaluable information to improve the efficiency of the gene-editing process so that we can hopefully correct disease-causing DNA mutations more quickly and precisely in the future,” said Professor Sriram Subramaniam, the study’s co-senior author.
Magnesium, required to activate the enzyme for DNA cleavage, was utilised for the process. Cas9 complexes were flash frozen and imaged using cryo-EM. The end results were snapshots of Cas9 trapped in three distinct structural arrangements while bound to nucleic acids. In two of the three states, the target DNA was cleaved but the enzyme remained bound to it.
In the first state, the enzyme was captured before cutting DNA. The second state imaged Cas9 almost immediately after the cut in the DNA was made. The enzyme’s active site was observed to hover above the cut in the DNA strand. In the third state, the enzyme began its movement toward its departure from the cleaved DNA.
“We discovered several new things about how Cas9 interacts with DNA and how it operates. One of the most interesting is how several enzyme domains move in concert and cycle between ordered and disordered states during reaction. This feature of the enzyme in which some domains are stable while others seem to be in a process of rapidly moving between different conformational states before settling down has not been seen nor predicted before,” said the study’s co-senior author, Miljan Simonovic. “This new information could be used to modulate how Cas9 processes targeted DNA and could help the design of better genome-editing tools.”
The findings were published in Nature Structural and Molecular Biology.
Related topics
CRISPR, Gene Therapy, Genome Editing, Genomics, Imaging, Microscopy
Related organisations
Nature Structural and Molecular Biology, University of British Columbia
Related people
Miljan Simonovic, Professor Sriram Subramaniam