Structure of commonly mutated histone chaperone revealed by new study
Posted: 10 March 2020 | Hannah Balfour (Drug Target Review) | No comments yet
Researchers have shown how ATAD2, a histone chaperone protein, may load histones on to DNA in order to create the chromatin structure.
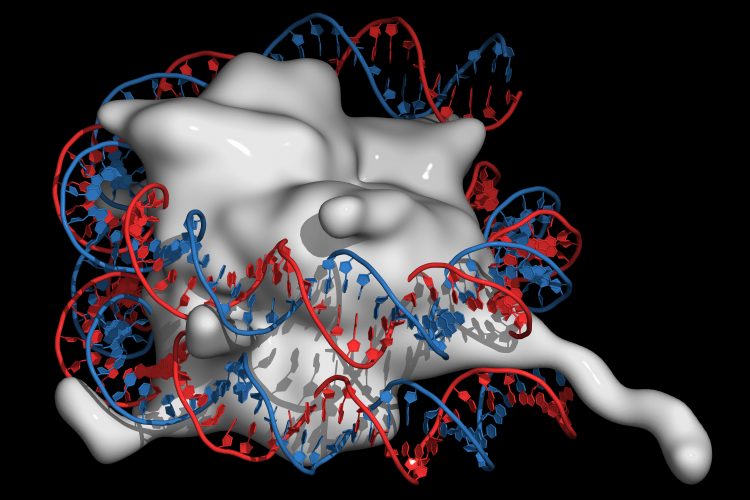

Researchers exploring how histones are loaded onto DNA and how the process may become misregulated have revealed the structure of a histone chaperone commonly mutated in cancers. The researchers hope the chaperone protein ATAD2 could now be targeted by novel cancer therapeutics.
Histones are loaded onto DNA to form the higher order structure known as chromatin. While histones primarily allow the large strands of DNA to fit inside the nucleus, they also protect the structure and sequence of DNA and control DNA replication and gene expression. Histone chaperones are proteins responsible for adding and removing specific histones onto DNA and are often mutated in cancers.
The paper, published in Nature Communications, focused on one histone chaperone, ATAD2 (also known as ANCCA), which is highly expressed in various cancers and associated with poor patient prognosis. ATAD2 is thought to regulate nucleosome density, the space between each group of eight histones wrapped in coils of DNA, by loading or removing a dimer of histones 3 and 4 (H3-H4). Nucleosome density decides if a protein will or will not be expressed: when nucleosomes are close together, the gene is not expressed; when they are spaced further apart, that segment of the DNA can be replicated and so the gene is expressed.
Using cryo-electron microscopy (Cryo-EM), a joint research team observed this chaperone protein in both native and near-native states, revealing the structural details of ATAD2. They presented cryo-EM structures of an ATAD2 family ATPase, Abo1, to atomic resolution in three different nucleotide states, showing unique structural features required for histone loading on DNA.

Three nucleosomes, each formed of eight histone proteins wrapped in DNA, on a single DNA strand.
They also used high-speed atomic force microscopy (HS-AFM) to visualise the transitions of Abo1 from an asymmetric spiral (ATP-state) to a symmetric ring (ADP- and apo-states). According to the researchers ATP-Abo1 likely binds histone tails and may facilitate the loading of H3-H4 onto DNA using energy from the release of a phosphate from ATP, leaving Abo1 in the ADP-state.
“This study is meaningful, as it reveals the structure and mechanism of histone chaperone proteins through the use of cutting-edge techniques in biophysical research, such as Cryo-EM,” said team co-leader, Professor Ja Yil Lee from the School of Life Sciences at Ulsan National Institute of Science and Technology (UNIST), South Korea. “This will accelerate the development of drug candidates, targeting ATAD2.”
The other institute involved in the study was the Department of Biological Sciences at the Korea Advanced Institute of Science and Technology (KAIST).
Related topics
Disease Research, DNA, Drug Targets, Histones, Imaging, Protein
Related conditions
Cancer
Related organisations
Korea Advanced Institute of Science and Technology, Ulsan National Institute of Science and Technology (UNIST)
Related people
Professor Ja Yil Lee