Study reveals how SARS-CoV-2 rewires human proteins for replication
Posted: 1 July 2020 | Victoria Rees (Drug Target Review) | No comments yet
A team of researchers has analysed how SARS-CoV-2 hijacks the proteins in its host cells to aid replication and infect nearby cells.

A team of researchers has analysed how SARS-CoV-2, the virus that causes COVID-19, hijacks the proteins in its target cells. The findings show how the virus shifts the cell’s activity to promote its own replication and to infect nearby cells. The scientists also identified seven clinically approved drugs that could disrupt these mechanisms and recommend that these drugs are immediately tested in clinical trials.
The collaboration included researchers at EMBL’s European Bioinformatics Institute (EMBL-EBI), the University of California San Francisco (UCSF), US, the Howard Hughes Medical Institute, US, the Institut Pasteur, France, and the University of Freiburg, Germany.
Viruses are unable to replicate and spread on their own so use a host to carry, replicate and transmit them to further hosts. To facilitate this process, viruses need to take control of their host cell’s machinery and manipulate it to produce new viral particles. Sometimes, this hijacking interferes with the activity of the host’s enzymes and other proteins.
“The virus prevents human cells from dividing, maintaining them at a particular point in the cell cycle. This provides the virus with a relatively stable and adequate environment to keep replicating,” said Pedro Beltrao, Group Leader at EMBL-EBI.
The scientists used mass spectrometry to evaluate all host and viral proteins that showed changes in phosphorylation (the addition of a phosphoryl group to a protein by a type of enzyme called a kinase) after SARS-CoV-2 infection. They found that 12 percent of the host proteins that interact with the virus were modified. The researchers also identified the kinases that are most likely to regulate these modifications.
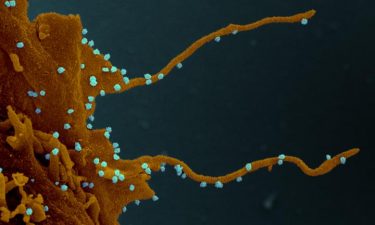

SARS-CoV-2 viruses visible on a cell with filopodia [credit: Elizabeth Fischer, Miscroscopy Unit NIH/NIAID].
One of the key findings from the study is that SARS-CoV-2-infected cells exhibit long, branched, arm-like extensions, or filopodia. According to the researchers, these structures may help the virus reach nearby cells in the body and advance the infection, but further study is warranted.
“The distinct visualisation of the extensive branching of the filopodia once again elucidates how understanding the biology of virus-host interaction can illuminate possible points of intervention in the disease,” said Nevan Krogan, Director of the Quantitative Biosciences Institute at UCSF.
“Kinases possess certain structural features that make them good drug targets. Drugs have already been developed to target some of the kinases we identified, so we urge clinical researchers to test the antiviral effects of these drugs in their trials,” said Beltrao.
In some patients, SARS-CoV-2 causes an overreaction of the immune system, leading to inflammation. An ideal treatment would relieve these exaggerated inflammatory symptoms while stopping the replication of the virus. Existing drugs targeting the activity of kinases may be the solution to both problems, say the researchers.
The team identified dozens of drugs approved by the Food and Drug Administration (FDA) or ongoing clinical trials that target the kinases of interest. Seven of these compounds, primarily anticancer and inflammatory disease compounds, demonstrated potent antiviral activity in laboratory experiments.
“Our data-driven approach for drug discovery has identified a new set of drugs that have great potential to fight COVID-19, either by themselves or in combination with other drugs and we are excited to see if they will help end this pandemic,” said Krogan.
The study was published in Cell.
Related topics
Disease Research, Drug Discovery, Drug Targets, Proteomics, Research & Development, Therapeutics
Related conditions
Coronavirus, Covid-19
Related organisations
EMBL European Bioinformatics Institute (EMBL-EBI), Howard Hughes Medical Institute, Institut Pasteur, University of California San Francisco (UCSF), University of Freiburg
Related people
Nevan Krogan, Pedro Beltrao