Computational modelling reveals SARS-CoV-2 mutation ‘hotspots’
Posted: 20 May 2021 | Victoria Rees (Drug Target Review) | No comments yet
A new modelling method could be used as a surveillance tool to monitor emerging infectious SARS-CoV-2 variants, scientists say.
A new study from New York University, US, and NYU Abu Dhabi, UAE, has used computational modelling to assess the biological significance of SARS-CoV-2 Spike (S) protein mutations, uncovering versions of the virus that bind more tightly or resist antibodies – offering a promising public health surveillance tool.
According to the researchers, new and more transmissible COVID-19 variants have emerged in recent months, fuelling surges of cases. As a public health measure, rapid surveillance methods are needed to determine the biological effects of variants and to help anticipate emerging infectious viral strains. However, monitoring new variants is no small task; genome sequencing shows that the SARS-CoV-2 S protein alone, for example, has about 5,000 possible variants.
“Screening such a large set of variants poses a tremendous challenge for conventional experimental methods,” said Hin Hark Gan, a senior research scientist at NYU and the study’s lead author. “An advantage of computer-based modelling is that a hundred mutations can be readily assessed in a few days.”
The scientists turned to a computational method that models how the S protein recognises the angiotensin-converting enzyme 2 (ACE2) receptor – a protein on the surface of many types of cells – to gain entry into host cells. Studies of coronaviruses indicate that S-ACE2 recognition is the basis for infection.
The researchers focused on screening the mutations located where the S protein and ACE2 receptor meet. They assessed 1,003 mutation combinations in the S and ACE2 proteins, including those resulting in the fast-spreading S variants that have originated in Brazil, South Africa, the UK and India.
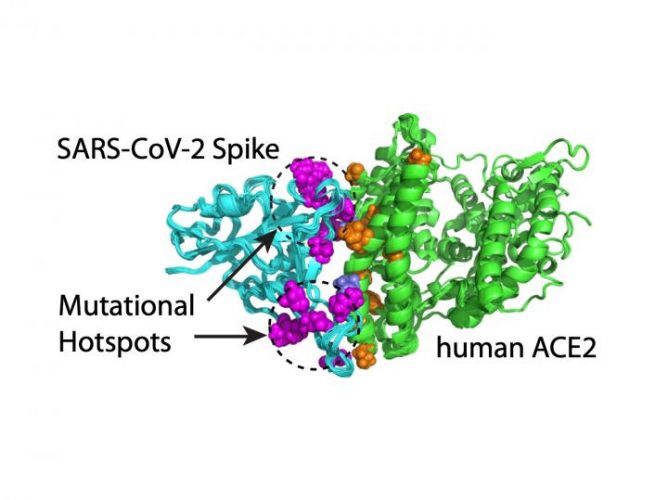
Computational modelling shows that mutations on SARS-CoV-2’s S protein that enhance the virus’ ability to bind to the ACE2 receptor occur in two clusters or mutation hotspots [credit: Hin Hark Gan and Kristin Gunsalus, NYU’s Department of Biology].
Their systematic assessment of variants revealed that S mutations which bind tightly to the ACE2 receptor occur in two clusters or mutation ‘hotspots’ on the binding interface. These hotspots are located in structurally flexible regions, indicating that mutations that increase binding effectively reprogrammed the spike conformation to enhance its recognition of the ACE2 receptor.
The researchers also looked at single, double and triple mutations in the critical S protein interface region, which make up some of the recently emerged infectious variants. Their analysis suggests the S variants S477N, N501Y and S477N + E484K and E484K + N501Y – fast-spreading double mutants found in Brazil, South Africa, US and UK – have increased binding to the ACE2 receptor relative to the original coronavirus that emerged in Wuhan.
The researchers observed that the SARS-CoV-2 E484K and E484Q mutations found in some recent fast-spreading variants are not only predicted to bind more strongly to ACE2 but have also been shown to confer antibody resistance. Neutralising antibodies are produced in response to viral infection and target different sites on the S protein to prevent the virus from invading host cells. This prompted the researchers to look at another factor contributing to viral transmission: antibody resistance of individual S mutations.
In particular, the variant circulating in India has two mutations in the S interface region: L452R and E484Q. This variant is not predicted to bind to the ACE2 receptor more tightly than the original virus, likely because the individual mutations have opposing effects (the L452R mutation binds less easily while the E484Q mutation binds more easily). However, both of these mutations are strong antibody evaders, a scenario not found in other recent variants.
“As more of antibody target sites become resistant to antibodies due to viral mutations, the efficacy of existing antibodies and vaccines may diminish,” said Gan. “This scenario is a likely cause for the rapid spread of the variant in India.”
The study not only provides explanations for the coronavirus’ rapid spread – both mutations that enhance binding to human cells and help evade antibodies – but also points to a promising predictive tool in the ongoing public health fight against SARS-CoV-2.
“Our computational modelling method can be used as a real-time surveillance tool to screen for emerging infectious COVID-19 variants. It allows for a more timely response to emerging outbreaks and could be used to guide the development of new vaccines,” said Professor Kristin Gunsalus, from NYU and the study’s senior author.
The study is published in the Journal of Molecular Biology.
Related topics
Antibodies, Bioinformatics, Disease Research, Informatics, Protein, Proteomics
Related conditions
Covid-19
Related organisations
New York University, NYU Abu Dhabi
Related people
Hin Hark Gan, Professor Kristin Gunsalus