CRISPR-based droplet method speeds up gene therapy research
Posted: 28 September 2021 | Anna Begley (Drug Target Review) | No comments yet
The new CRISPR-based technology called MIC-Drop rapidly identified several genes for heart development and function in zebrafish.
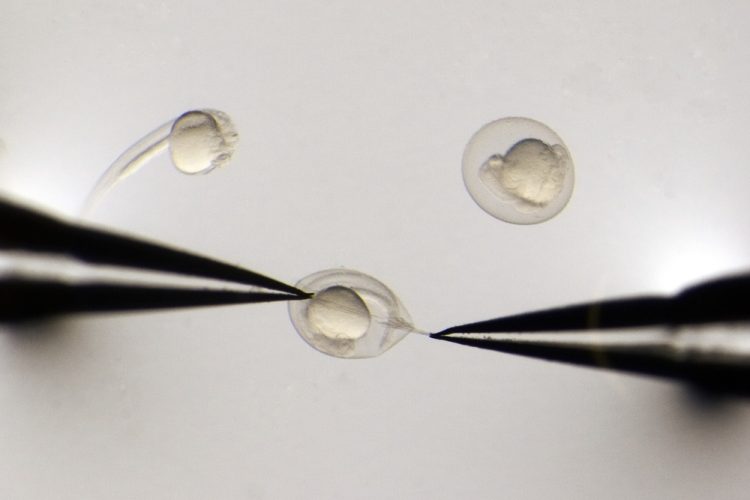

Researchers at the University of Utah, US, have developed a technology that enables scientists to deploy the CRISPR gene editing system into zebrafish to rapidly evaluate the functions of hundreds of genes in a single experiment. The advance marks the first time that screens using the robust, Nobel-prize winning CRISPR technology have been possible in any animal model.
Previously, setting up a CRISPR screen of hundreds of genes in zebrafish would have taken a team of researchers many days and required hundreds of needles, commented researcher Saba Parvez. Usually, to inactivate a single gene in a zebrafish embryo, scientists prepare a guide RNA targeting that gene, then mix it with the Cas9 enzyme, load the solution into a needle and inject a carefully calibrated volume of the solution into the embryo. If they want to inactivate a different gene in a different embryo, they must load a new needle with a new Cas9-guide RNA solution. “The process has always been focused on a single gene or a single modification at a time,” summarised Randall Peterson, who led the study. “So if you want to do 100 genes, it is 100 times as much work.”
NEW ARTICLE – CLICK BELOW TO READ:
CRISPR-Cas9-based gene dropout screens: a powerful platform for drug discovery
MIC-Drop, which stands for Multiplexed Intermixed CRISPR Droplets, solves that problem by packaging the components of the CRISPR system into microscopic oil-encased droplets, which can mingle together without mixing up their contents. To set up a screen of many genes with MIC-Drop, the researchers began by creating a library of guide RNAs. Each guide RNA is packaged into its own droplet, along with the Cas9 enzyme. To keep track of target genes, every droplet also includes a DNA barcode identifying its contents.
The team fine-tuned the chemistry of the droplets to ensure they would remain stable and discrete, so droplets designed to target different genes can be mixed together and loaded into the same needle. Under a microscope, the MIC-Drop user injects a single droplet into a zebrafish embryo, then moves on to the next embryo and injects the next droplet. The process can be repeated hundreds of times, delivering a single packet of CRISPR components to each embryo, so that in every embryo, the system inactivates a single gene. The researchers then monitor the animals for potential effects.
To demonstrate the potential of MIC-Drop, the team tested 188 different zebrafish genes for a potential role in heart development. After creating guide RNAs targeting those genes and introducing the CRISPR system into hundreds of fish embryos, they identified several animals that developed heart defects as they matured. Using the DNA barcodes in those fish, the team were able to trace the defects back to 13 different inactivated genes. Due to the similarities between zebrafish and human genes, the finding may point toward previously unknown aspects of heart development in humans.
The study was published in Science.
Related topics
CRISPR, DNA, Enzymes, Gene Therapy, Genetic Analysis, Genomics, Technology
Related organisations
University of Utah
Related people
Randall Peterson, Saba Parvez