Discovering how drugs combat HIV and why resistance can occur
Posted: 17 February 2020 | Victoria Rees (Drug Target Review) | No comments yet
A group of researchers, led by Professor Zucai Suo, have revealed the mode of action of two HIV drugs and identified how resistance can develop, which they say could lead to improved drug design in the future.

What can researchers gain from understanding the molecular interactions of drugs that patients frequently develop resistance to? According to a recent study, where US researchers discovered the mode of action of two widely used HIV antiviral drugs, their findings will improve the design of future therapeutics, making them even more effective and prevent the spread of resistance.
Identifying the mechanism
The team of researchers from Florida State University (FSU) College of Medicine established the previously unknown workings of two antiviral drugs used to treat patients with HIV, outlined in their paper published in Communications Biology.
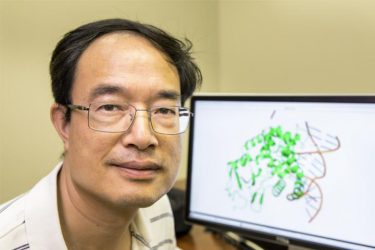


Professor Zucai Suo of the FSU College of Medicine. His research has established the mechanism responsible for how two widely used antiviral drugs inhibit viruses, which could create more treatment options for patients with HIV or hepatitis B (credit: Colin Hackley/Florida State University College of Medicine).
Zucai Suo, Eminent Professor and the Dorian and John Blackmon Chair in Biomedical Science at the FSU College of Medicine, and his colleagues investigated how a single HIV-1 mutation can inactivate the anti-HIV drugs emtricitabine and lamivudine. The former is also approved for the treatment of hepatitis B.
Based on their findings, the researchers have proposed new pathways for developing HIV drugs that can avoid viral mutations that render certain treatments ineffective for many patients.
Potent analogue drugs
The researchers’ paper suggests new chemical structures for more potent L-nucleoside analogue drugs, which may possess different drug-resistance mutation profiles from the most widely used current anti-HIV drugs.
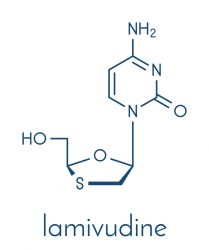

The researchers explain that once the drugs are administered, they are absorbed into cells and converted by kinases into active triphosphate metabolites. This form is recognised by HIV-1 RT and incorporated into a DNA primer strand, preventing viral replication.
Although this was already understood, the drugs were discovered through blind trials. Important details about the underlying mechanism triggering L-NRTIs have therefore remained a mystery.
Suo, the study’s co-lead author, discussed with Drug Target Review how the group established the mechanisms behind the antiviral drugs. By employing both enzyme kinetics and X-ray crystallography, they were able to fully understand the inhibitory mechanism of the L-nucleoside drugs on HIV-1 reverse transcriptase.
They found that the RT enzyme has a unique pocket and supposedly recognises nucleoside reverse transcriptase inhibitors (NRTIs), but not their structural mirror images, L-NRTIs.
Although the researchers did not investigate how these drugs inhibit hepatitis B polymerase in this study, Suo said they believe the mechanism will be similar, and therefore useful for this condition as well.
Resistance to therapy
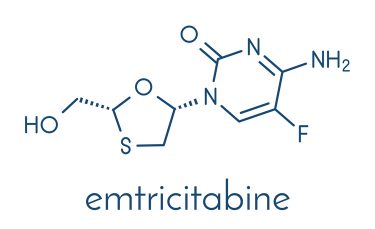

In the case of emtricitabine and lamivudine, M184V produces resistance by sterically hindering the drugs’ oxathiolane ring and forcing phosphates from the DNA primer strand, which prevents them from acting against HIV.
Based on the structures defined in their paper, the team suggests that that other L-nucleoside analogue drugs could be developed by modifying atoms in the nucleobase. Suo hopes that some will be more potent and possess different mutation profiles relative to the current drugs emtricitabine and lamivudine.
In their paper, the researchers propose strategies to combat resistance:
- adjusting the steric bulk around the central ring to prevent phosphates from flipping to a non-productive state, which would only allow the phosphates to exist in a productive conformation. This could potentially be achieved with substitutions to the oxathiolane ring which strengthens it but would still be accepted in the binding pocket
- optimisation of NRTI interactions with R72, part of the phosphate side chain, could increase potency against HIV-1 RT as this residue is essential for polymerase function. Identifying similar residues to R72 in other viral polymerases could also be a potential option for improving the efficacy of drug therapies against viruses.
Suo said that it may be best to “design derivatives of emtricitabine and lamivudine in order to avoid similar mutations,” allowing for a higher number of patients to be treated.
The significance of the findings
Suo explained that previously, drugs in this class were discovered by random screening, but the exact inhibitory mechanism was unknown. However, now that the researchers have elucidated how emtricitabine and lamivudine work, scientists will be able to build on this mechanism and make necessary structural changes to make better, or more powerful, new drugs.
Designing drugs for the future
In their paper, the researchers note that these comprehensive kinetic and structural studies provide the most definitive insights into the binding modes of oxathiolane analogues and the mechanism of NRTI resistance achieved by the M184V mutation of HIV-1 RT.

He said that these new drugs are important when considering the fact that each current antiviral drug causes a unique drug-resistant mutation profile in clinics and that more treatment options are needed to counter drug resistance.
Related topics
Antiretroviral Therapies, Disease Research, Drug Development, Drug Targets, Therapeutics
Related conditions
Hepatitis B, HIV
Related organisations
Florida State University (FSU) College of Medicine
Related people
Professor Zucai Suo