Genomics to match treatments to pancreatic cancer patients
Posted: 4 March 2019 | Nic Losardo (Drug Target Review) | No comments yet
Genomics could be used to match drugs to patients better, with these genetic signatures also giving an earlier insight into detection of the condition…
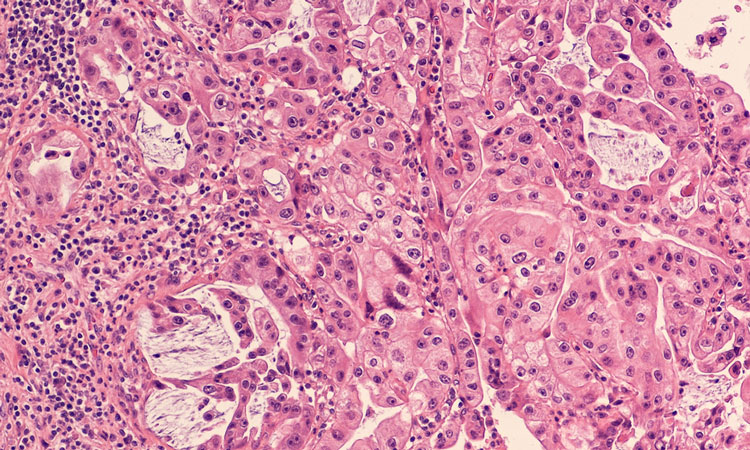
Pancreatic cancer is a grim diagnosis, with a five-year survival rate of less than 9 percent. To improve these odds, researchers at UPMC and the University of Pittsburgh School of Medicine sought genetic signatures in the largest study of its kind that could be used to better match drugs to patients and for early detection.
The study involved sifting through the genomes of thousands of tumors, sampled from all over the world. In 17 percent of cases, there was a genetic flag that indicated the tumor should be susceptible to existing chemotherapy drugs. The researchers also found supporting evidence for heritable genes, including some in the BRCA family associated with breast cancer, that can predispose whole families toward pancreatic cancer.
“People have been looking for such markers for a long time, and our study shows that it’s possible to break pancreatic cancer patients into different treatment buckets,” said senior author Dr Nathan Bahary, Oncologist at UPMC Hillman Cancer Center and Associate Professor of Medicine at Pitt.
One reason why pancreatic cancer is so deadly is that the majority of patients often are identified late in their disease course and frequently present with inoperable tumors at the time of diagnosis. For some of these patients, it may be possible to shrink the tumor with existing chemotherapy drugs, but in a disease where 75 percent of patients die within a year of diagnosis, time is of the essence, and unfortunately, there’s no way to know in advance which patients will respond to which drugs.
“Every pancreatic cancer is different, and performing molecular profiling of each patient’s tumor could help determine the best treatment options,” said lead author Dr Aatur Singhi, surgical pathologist at UPMC and Assistant Professor of Pathology at Pitt. “Rather than blindly giving patients the same chemotherapy, we want to tailor a patient’s chemo to their tumor type. A one-size-fits-all approach isn’t going to work. Therefore, we would like to make molecular profiling standard-of-care for patients with pancreatic cancer.”
Prof Singhi and Prof Bahary’s study characterised the genome of 3,594 pancreatic tumor samples from patients around the world, provided by collaborators at Foundation Medicine.
“We believe that this is the largest study in pancreatic cancer conducted using comprehensive genomic profiling to identify a broad set of genomic alterations, and ultimately, therapeutic targets, in this difficult-to-treat disease,” said Dr Siraj Ali, Senior Director of Clinical Development at Foundation Medicine.
Besides shrinking tumors with personalised chemotherapy, another way to increase pancreatic cancer survival rates is through increased pancreatic cysts screening, Prof Singhi said, but the problem is that pancreatic cysts are incredibly common, and not all lead to cancer.
Previously, Prof Singhi and colleagues developed a clinical molecular test known as PancreaSeq to evaluate common pancreatic cysts and identify which cases may progress to cancer. The discovered biomarkers can be added to the PancreaSeq platform, already being used by several institutions, including UPMC.
The study was published in the journal Gastroenterology.
Related topics
Disease Research, Drug Discovery, Genomics, Oncology, Research & Development, Therapeutics
Related conditions
Pancreatic cancer
Related organisations
University of Pittsburgh School of Medicine, UPMC
Related people
Dr Aatur Singhi, Dr Nathan Bahary, Dr Siraj Ali