Crystals mapped to advance treatments for stroke, diabetes and dementia
Posted: 5 July 2019 | Drug Target Review | No comments yet
Researchers haved mapped the crystal structure of a protein to find out how a drug latches onto it.
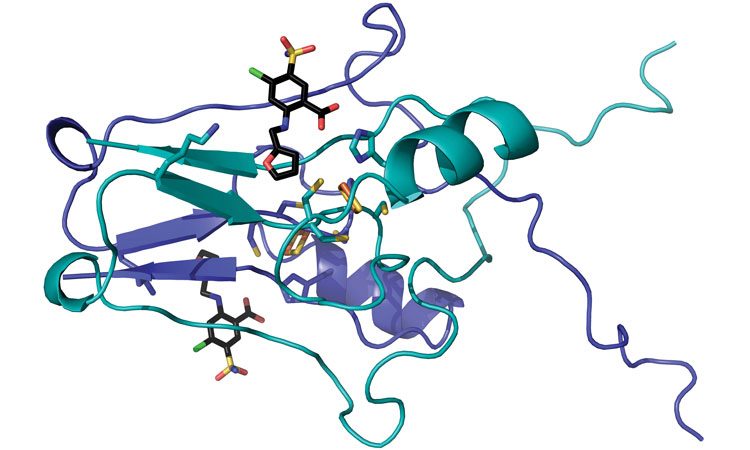

A team of WVU researchers (including Werner Geldenhuys, John Hollander and Aaron Robart) have mapped the crystal structure of a protein called “mitoNEET” and pinpointed how a drug latches on it. Because previous research has implicated mitoNEET in a diabetes, stroke and heart disease, the researchers’ findings may inform the development of new treatments for those tough-to-tackle conditions (credit: WVU).
Researchers at West Virginia University have mapped the crystal structure of a protein that resides in our cells and for the first time determined how a drug latches onto it.
The study centered on the protein, MitoNEET, which inhabits the outer membrane of our mitochondria and energises cells.
“MitoNEET is a novel therapeutic target for metabolic-based diseases and could possibly lead to disease-modifying treatments for Alzheimer’s disease and stroke,” said Werner Geldenhuys, an associate professor in the School of Pharmacy and School of Medicine. He and his colleagues (including Aaron Robart, an assistant professor in the WVU School of Medicine, John Hollander, assistant dean for professional programs in the WVU School of Medicine, and Timothy Long, an associate professor in the Marshall University School of Pharmacy) carried out the project.
“This protein has been implicated in a lot of diseases that are very tough to tackle: things like diabetes, stroke, heart disease,” added Robart.
The researchers isolated mitoNEET from both bacterial overexpression and animal models and then synthesised 11 molecules similar to furosemide (a common diuretic sold under the brand name LASIX) and exposed the mitoNEET to them.
After the molecules bonded to the mitoNEET, the researchers built atom-by-atom maps of the pairings. They remotely controlled Argonne National Laboratory’s Advanced Photon Source, which bombards samples with ultra-bright, high-energy X-rays, to reveal precisely how the molecules came together.
The team then discovered that the molecules docked into a cluster of iron and sulfur atoms that made up part of the protein.
“These findings are of importance as they allow us to continue to understand the role played by mitochondria and bioenergetics in many disease states,” Hollander said. “The modulation of mitochondrial function through targeted therapeutics may be a critical avenue of drug discovery.”
Understanding mitoNEET’s cellular function could improve the performance of drugs that work by altering the protein’s activity. For example, adding an extra oxygen group to a drug’s molecular structure could dramatically tighten its bond to mitoNEET and eliminate unintended binding to other cellular proteins.
“The success of this project really illustrates how approaches that are considered basic science can provide considerable insight into clinical problems,” commented Michael Schaller, who chairs the School of Medicine’s Department of Biochemistry.
The findings of this study appear in Communications Chemistry.
Related topics
Analysis, Disease Research, Drug Discovery, Protein, Research & Development, Targets, Therapeutics
Related conditions
Alzheimer's, Diabetes, Stoke
Related organisations
West Virginia University
Related people
Aaron Robart, John Hollander, Michael Schaller, Timothy Long, Werner Geldenhuys